Enalos Cloud Platform IDs two molecules that inhibit proteins involved in chronic inflammatory disease
Virtual drug screening approach could speed discovery of potential new treatments
Scientists have identified two small molecules that could be pursued as potential treatments for chronic inflammatory diseases. According to a paper published in PLOS Computational Biology, the researchers singled out the molecules using a new drug screening approach they developed.
Both molecules, known as T23 and T8, inhibit the function of a protein called tumor necrosis factor (TNF), which is involved in inflammation in diseases such as rheumatoid arthritis, Crohn's disease, psoriasis, multiple sclerosis, and more. Drugs that inhibit TNF's function are considered the most effective way to combat such diseases. However, not all patients respond to them, and their effectiveness can wear off over time.
To aid discovery of better TNF inhibitor drugs, Georgia Melagraki and colleagues from Greece and Cyprus developed a new computer-based drug screening platform. The platform incorporates proprietary molecular properties shared between TNF and another protein called RANKL, which is also involved in chronic inflammatory diseases.
The researchers developed the platform based on a combination of advanced computational tools. The platform was then used to virtually screen nearly 15,000 small molecules with unknown activity and to predict their interactions with the TNF and RANKL proteins; specifically, how well the small molecules might disrupt the protein-protein interactions (PPIs) leading to the trimerization and activation of these crucial proteins. "This virtual experiment identified nine promising molecules out of thousands of candidates," says study co-corresponding author Antreas Afantitis of NovaMechanics Ltd, Cyprus.
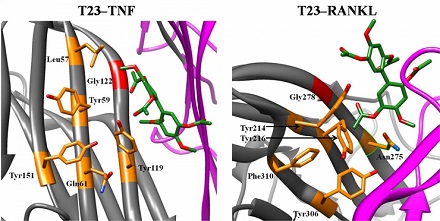
To further evaluate their potential, the scientists studied how the nine small molecules interacted with TNF and RANKL in real-world laboratory experiments. Of the nine molecules, T23 and T8 surfaced as particularly strong TNF inhibitors. Both molecules bind to TNF and RANKL, preventing them from interacting properly with other proteins. Both also show low potential for causing toxic side effects in humans.
With further research, T23 and T8 could be "further optimized to develop improved treatments for a range of inflammatory, autoimmune, and bone loss diseases," says study co-corresponding author George Kollias of the Biomedical Sciences Research Center 'Alexander Fleming', Greece. Meanwhile, the new virtual drug screening approach could enable discovery of other promising TNF inhibitors, and could be modified to search for potential treatments for additional diseases.
NovaMechanics has significant expertise on the development of software especially designed for the risk assessment, read across and grouping of small molecules and nanomaterials. NovaMechanics web services (Enalos Cloud Platform) are based solely on open source and in house software that were developed with the purpose of making in silico models available to the interested user wishing to generate evidence on potential biological effects in the decision making framework. These web services facilitate computer aided nanoparticles/ small molecules design as they can serve as a source of activity prediction for novel nano-structures and small molecules.
The Enalos Cloud Platform is a platform that hosts several predictive models for drug discovery and risk assessment of small molecules and nanoparticles. In the previous steps, they succeeded to develop a predictive consensus model for TNF inhibition. Since, to the best of their knowledge, this is the only ligand–based model developed from an extensive dataset of TNF inhibitors, they decided to release it as a web service to facilitate the need for the design and virtual screening of novel potent small–molecule inhibitors of TNF, by providing immediate access to the model’s results. The model can be easily accessed through http://enalos.insilicotox.com/TNFPubChem.
Access freely available article in PLOS Computational Biology: http://journals.
