Germany creates AI software for dependable imaging of proteins in cells
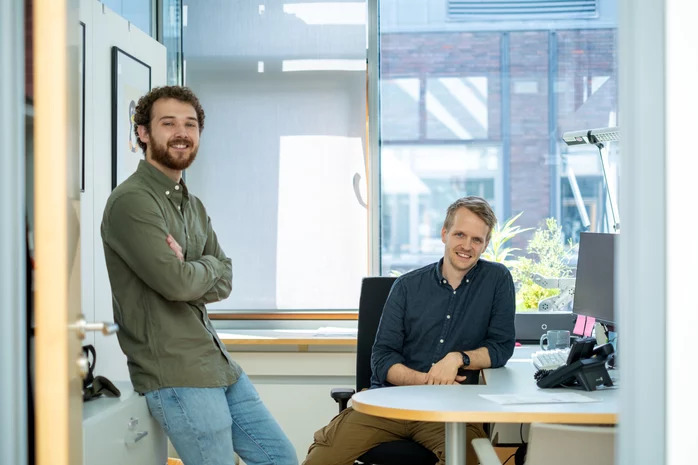

Max Planck researchers from Dortmund programmed a tool that accurately recognizes and picks proteins in electron cryo-tomography, substituting troublesome hand selection
Electron cryo-tomography (cryo-ET) is emerging as a powerful technique to provide detailed 3D images of cellular environments and enclosed biomolecules. However, one of the challenges of the methodology is the identification of protein molecules in the images for further processing. A research team around Stefan Raunser, Director at the MPI of Molecular Physiology in Dortmund, led by Thorsten Wagner, developed software to pick proteins in crowded cellular volumes. The new open-source tool, called TomoTwin, is based on deep metric learning and allows scientists to locate several proteins with high accuracy and throughput without manually creating or retraining the network each time. 
The more, the better
“TomoTwin paves the way for automated identification and localization of proteins directly in their cellular environment, expanding the potential of cryo-ET,” says Gavin Rice, co-first author of the publication. Cryo-ET has the potential to decipher how biomolecules work within a cell and, by that, to unveil the basis of life and the origin of diseases.
In a cryo-ET experiment, scientists use a transmission electron microscope to obtain 3D images, called tomograms, of the cellular volume containing complex biomolecules. To gain a more detailed image of each different protein, they average as many copies of them as possible – similar to photographers capturing the same photo at varying exposures to later combine them in a perfectly exposed image. Crucially, one has to correctly identify and locate the different proteins in the picture before averaging them. “Scientists can attain hundreds of tomograms per day, but we lacked tools to fully identify the molecules within them,” says Rice.
Hand-picking
So far, researchers used algorithms based on templates of already known molecular structures to search for matches in the tomograms, but these tend to be error-prone. Identifying molecules by hand is another option that ensures high-quality picking but takes days to weeks per dataset.
Another possibility would be to use a form of supervised machine learning. These tools can be very accurate but currently lack usability, as they require manually labeling thousands of examples to train the software for each new protein, an almost impossible task for small biological molecules in a crowded cellular environment.
TomoTwin
The newly developed software TomoTwin overcomes many of these obstacles: It learns to pick the molecules that are similar in shape within a tomogram and maps them to a geometric space – the system is rewarded for placing similar proteins near each other and penalized otherwise. In the new map (image 1), researchers can isolate and accurately identify the different proteins and use this to locate them inside the cell. “One advantage of TomoTwin is that we provide a pre-trained picking model,” says Rice. By removing the training step, the software can even run on local computers – where processing a tomogram usually requires 60-90 minutes, and runtime on the MPI supercomputer Raven is reduced to 15 minutes per tomogram.
TomoTwin allows researchers to pick dozens of tomograms in the time it takes to manually pick a single one, therefore increasing the throughput of data and the averaging rate to obtain a better image. The software can currently locate globular proteins or protein complexes larger than 150 kilodaltons in cells; in the future, the Raunser group aims to include membrane proteins, filamentous proteins, and proteins of smaller sizes.
POPULAR
- NASA predicts the Sun's corona behavior, revealing its mysteries using advanced computational methods
- Australia on track for unprecedented, decades-long megadroughts: Supercomputer modeling raises concern
- Revolutionizing precision agriculture: The impact of Transformer Deep Learning on water, energy demands

 How to resolve AdBlock issue?
How to resolve AdBlock issue?