Within the human body, the delivery of oxygen occurs at the microscale, where the disciplines of physics, chemistry, and biology intersect. Elucidating the mechanisms by which oxygen is transported through the bloodstream, diffuses out of erythrocytes, and is utilized by surrounding tissues has remained a formidable challenge, primarily due to the inherent complexity of these processes.
Recently, a study published in the International Journal of Heat and Mass Transfer introduced a significant advancement: a fully three-dimensional computational model that simultaneously simulates oxygen transport alongside the motion and deformation of individual red blood cells (RBCs). This development constitutes an important step toward addressing one of the most complex multiphysics challenges in biomedical science.
A problem too complex to see directly
At the scale of capillaries, oxygen transport is governed by a delicate interplay of mechanisms:
- Fluid flow through narrow vessels
- Diffusion across multiple regions (cells, plasma, tissue)
- Chemical reactions involving hemoglobin
- Continuous deformation and interaction of red blood cells
Traditional models simplified this system, often ignoring individual cells or treating vessels as static tubes. But such approximations fall short of capturing how oxygen is actually delivered in living tissue.
The new study breaks from that tradition by embracing the full complexity.
A unified multiphysics framework
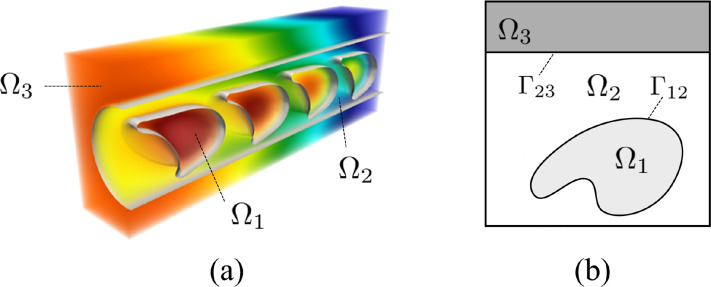
The researchers developed a diffuse interface model that unifies multiple physical processes into a single computational framework. Instead of treating boundaries, like the surface of a red blood cell, as sharp discontinuities, the method smooths them into a continuous transition region. This allows the governing equations to be solved seamlessly across the entire domain.
At its core, the model simultaneously solves:
- The incompressible Navier–Stokes equations for blood flow
- Advection–diffusion–reaction equations for oxygen transport
- Fluid–structure interaction governing deformable red blood cells
Red blood cells are modeled as elastic membranes interacting with fluid using an immersed boundary method, enabling them to move, deform, and respond dynamically to their environment.
The result is a fully coupled 3D simulation where flow, chemistry, and cellular mechanics evolve together.
Capturing the behavior of living blood
One of the most striking outcomes of the study is the ability to simulate how red blood cells actively regulate oxygen delivery.
Rather than acting as passive carriers, the simulations suggest that RBCs:
- Adjust oxygen release based on local tissue demand.
- Interact with one another in ways that influence flow distribution.
- Contribute to maintaining relatively uniform oxygenation across tissue.
This emergent behavior, arising purely from physics and chemistry, offers new insight into how the body maintains balance at the microscale.
The computational challenge beneath the surface
While the study does not explicitly reference supercomputers or high-performance computing (HPC) systems, the scale and sophistication of the model place it firmly within the realm of HPC-class workloads.
The simulation involves:
- Three-dimensional, time-dependent PDEs
- Moving and deforming interfaces
- High-order numerical schemes (including fifth-order advection methods)
- Coupled nonlinear physics across multiple domains
These are precisely the kinds of problems that increasingly drive demand for advanced computing infrastructure.
Interestingly, rather than relying solely on brute-force computational power, the researchers focused on algorithmic efficiency:
- A mixture formulation eliminates the need for complex interface reconstruction.
- Fixed Cartesian grids simplify geometry handling.
- Carefully chosen numerical schemes balance accuracy and cost.
This approach reflects a broader trend in computational science: pairing smarter algorithms with scalable hardware to tackle previously intractable problems.
A glimpse of scalable biomedical simulation
The implications extend beyond this specific study. By demonstrating a practical way to simulate oxygen transport with deformable cells in 3D, the work lays a foundation for:
- Patient-specific microcirculation modeling
- Disease studies involving impaired oxygen delivery
- Integration with larger-scale physiological simulations
As these models grow in size and realism, they are likely to transition naturally onto parallel and high-performance computing platforms, where their full potential can be realized.
Looking Ahead
This research highlights a subtle but important shift. The frontier of biomedical modeling is no longer defined solely by biological insight, but increasingly by computational capability.
Even when supercomputers are not explicitly named, they linger in the background, implicit in the complexity of the equations, the dimensionality of the models, and the ambition of the questions being asked.
In that sense, this study is not just about oxygen transport. It is a preview of a future where understanding life at its smallest scales depends as much on computational innovation as it does on biology itself.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?