A new study highlights a pivotal shift in biomedical research: breakthroughs now depend as much on powerful computational tools as on laboratory instruments. Driving this transformation is the RAMSES supercomputer at the University of Cologne's IT Center in Germany, empowering researchers to process and analyze enormous digital pathology datasets at a scale previously unattainable.
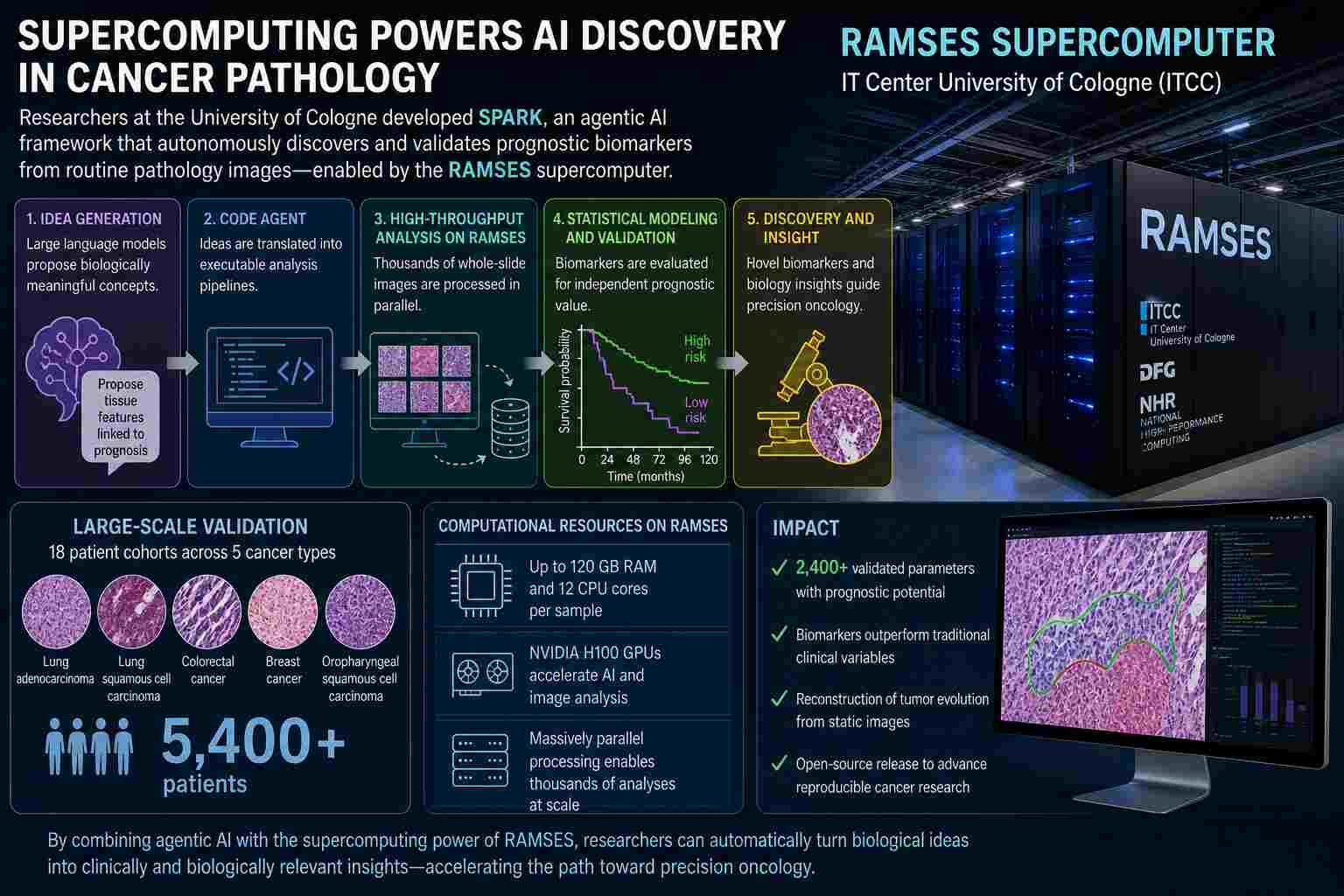
Featured in a recent Nature Medicine publication, this work presents SPARK, an advanced AI-driven framework described as “agentic” for its ability to autonomously generate, test, and validate hypotheses in cancer pathology. While the conceptual innovation is noteworthy, it is RAMSES’s computational power that makes such a system practically feasible.
Scaling digital pathology beyond human limits
Digital pathology operates on whole-slide images (WSIs), each of which can reach gigapixel resolution. When multiplied across thousands of patient samples and multiple cancer types, the resulting data volume quickly becomes prohibitive for conventional computing.
To address this, researchers deployed a hybrid computational architecture in which high-throughput workloads were executed on RAMSES, a high-performance computing (HPC) system designed for large-scale modeling and simulation. The system integrates advanced GPU resources, including multiple NVIDIA H100 accelerators, optimized for parallel processing of AI and image analysis pipelines.
Within this environment, each pathology case required substantial dedicated resources, up to 120 GB of memory and 12 CPU cores per sample, highlighting the intensity of the computational workload.
The SPARK framework: AI at scale
The SPARK system represents a shift from static machine learning models to dynamic, reasoning-based AI workflows. Rather than being trained solely on labeled data, SPARK generates its own analytical “ideas,” translates them into executable code, and evaluates their predictive value across large datasets.
This process unfolds in several stages:
- Idea generation using large language models (LLMs)
- Automated code creation and validation
- High-throughput parameter extraction from WSIs
- Statistical modeling and prognostic evaluation
While early-stage development and prototyping could be conducted on smaller systems, the full-scale execution, particularly across cohorts exceeding 5,000 patients, required the parallel processing capabilities of RAMSES.
High-performance computing meets oncology
The integration of HPC into this workflow enabled several key advances:
1. Massive Parallel Image Analysis
RAMSES allowed simultaneous processing of thousands of WSIs, performing segmentation, cell classification, and spatial mapping across seven distinct cell types.
RAMSES allowed simultaneous processing of thousands of WSIs, performing segmentation, cell classification, and spatial mapping across seven distinct cell types.
2. Large-Scale Parameter Exploration
The system generated and evaluated thousands of candidate biomarkers, over 2,400 validated parameters in some analyses, each representing a potential predictor of cancer progression.
The system generated and evaluated thousands of candidate biomarkers, over 2,400 validated parameters in some analyses, each representing a potential predictor of cancer progression.
3. Predictive Modeling at Population Scale
Using HPC resources, the team conducted multivariable statistical modeling across diverse cancer cohorts, identifying features with independent prognostic value beyond traditional clinical metrics.
Using HPC resources, the team conducted multivariable statistical modeling across diverse cancer cohorts, identifying features with independent prognostic value beyond traditional clinical metrics.
4. Temporal Reconstruction of Tumor Evolution
By analyzing spatial patterns within tumors, the system inferred evolutionary sequences of disease progression, an inherently data-intensive task requiring both computational power and algorithmic sophistication.
By analyzing spatial patterns within tumors, the system inferred evolutionary sequences of disease progression, an inherently data-intensive task requiring both computational power and algorithmic sophistication.
RAMSES: Infrastructure as an enabler of discovery
The RAMSES system, formally known as the Research Accelerator for Modeling and Simulation with Enhanced Security, played a central role in enabling these analyses. Hosted at the University Hospital Cologne and supported by national and European funding initiatives, it provides a secure, scalable environment for data-intensive biomedical research.
Crucially, RAMSES is not merely a computing resource but an integrated platform supporting:
- GPU-accelerated AI workloads
- High-memory nodes for large dataset handling
- Parallelized pipelines for image and statistical analysis
- Secure processing of sensitive clinical data
Without such infrastructure, the SPARK framework would be constrained to small-scale experiments rather than clinically relevant population studies.
Toward autonomous scientific discovery
The implications of this work extend beyond pathology. By combining agent-based AI systems with supercomputing infrastructure, researchers are moving toward autonomous scientific discovery pipelines, systems that can generate hypotheses, test them, and refine their own analytical strategies.
In oncology, this approach could accelerate the identification of novel biomarkers, improve patient stratification, and ultimately inform more personalized treatment strategies. More broadly, it signals a shift in how science is conducted: from manually driven analysis to computational ecosystems capable of operating at scale.
The supercomputing imperative in modern medicine
The study reinforces a central theme in contemporary research: data alone is not enough. The ability to extract meaning from complex, high-dimensional datasets depends critically on access to advanced computational infrastructure.
In this case, the RAMSES supercomputer transformed a conceptual AI framework into a practical, high-impact tool, demonstrating that in the era of digital medicine, supercomputing is not an accessory but a necessity.
As biomedical datasets continue to expand in size and complexity, systems like RAMSES will increasingly define the boundary between theoretical possibility and real-world application.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?