The European Southern Observatory’s Very Large Telescope Interferometer (ESO’s VLTI) has obtained the deepest and sharpest images to date of the region around the supermassive black hole at the center of our galaxy. The new images zoom in 20 times more than what was possible before the VLTI and have helped astronomers find a never-before-seen star close to the black hole. By tracking the orbits of stars at the center of our Milky Way, the team has made the most precise measurement yet of the black hole’s mass. 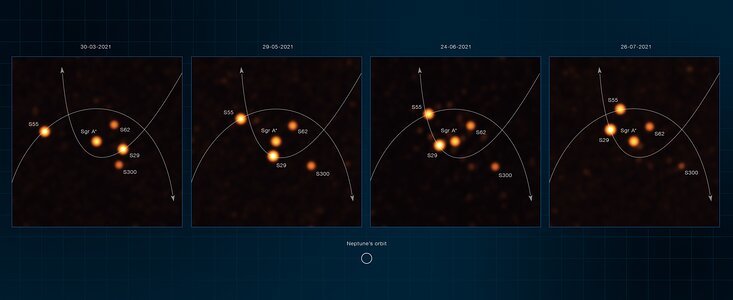
“We want to learn more about the black hole at the center of the Milky Way, Sagittarius A*: How massive is it exactly? Does it rotate? Do stars around it behave exactly as we expect from Einstein’s general theory of relativity? The best way to answer these questions is to follow stars on orbits close to the supermassive black hole. And here we demonstrate that we can do that to a higher precision than ever before,” explains Reinhard Genzel, a director at the Max Planck Institute for Extraterrestrial Physics (MPE) in Garching, Germany who was awarded a Nobel Prize in 2020 for Sagittarius A* research. Genzel and his team’s latest results, which expand on their three-decade-long study of stars orbiting the Milky Way's supermassive black hole, are published today in two papers in Astronomy & Astrophysics.
On a quest to find even more stars close to the black hole, the team, known as the GRAVITY collaboration, developed a new analysis technique that has allowed them to obtain the deepest and sharpest images yet of our Galactic Centre. “The VLTI gives us this incredible spatial resolution and with the new images, we reach deeper than ever before. We are stunned by their amount of detail, and by the action and number of stars they reveal around the black hole,” explains Julia Stadler, a researcher at the Max Planck Institute for Astrophysics in Garching who led the team’s imaging efforts during her time at MPE. Remarkably, they found a star, called S300, which had not been seen previously, showing how powerful this method is when it comes to spotting very faint objects close to Sagittarius A*.
With their latest observations, conducted between March and July 2021, the team focused on making precise measurements of stars as they approached the black hole. This includes the record-holder star S29, which made its nearest approach to the black hole in late May 2021. It passed it at a distance of just 13 billion kilometers, about 90 times the Sun-Earth distance, at the stunning speed of 8740 kilometers per second. No other star has ever been observed to pass that close to, or travel that fast around, the black hole.
The team’s measurements and images were made possible thanks to GRAVITY, a unique instrument that the collaboration developed for ESO’s VLTI, located in Chile. GRAVITY combines the light of all four 8.2-meter telescopes of ESO’s Very Large Telescope (VLT) using a technique called interferometry. This technique is complex, “but in the end you arrive at images 20 times sharper than those from the individual VLT telescopes alone, revealing the secrets of the Galactic Centre,” says Frank Eisenhauer from MPE, principal investigator of GRAVITY.
“Following stars on close orbits around Sagittarius A* allows us to precisely probe the gravitational field around the closest massive black hole to Earth, to test General Relativity, and to determine the properties of the black hole,” explains Genzel. The new observations, combined with the team’s previous data, confirm that the stars follow paths exactly as predicted by General Relativity for objects moving around a black hole of mass 4.30 million times that of the Sun. This is the most precise estimate of the mass of the Milky Way’s central black hole to date. The researchers also managed to fine-tune the distance to Sagittarius A*, finding it to be 27 000 light-years away.
To obtain the new images, the astronomers used a machine-learning technique, called Information Field Theory. They made a model of how the real sources may look, simulated how GRAVITY would see them, and compared this simulation with GRAVITY observations. This allowed them to find and track stars around Sagittarius A* with unparalleled depth and accuracy. In addition to the GRAVITY observations, the team also used data from NACO and SINFONI, two former VLT instruments, as well as measurements from the Keck Observatory and NOIRLab’s Gemini Observatory in the US.
GRAVITY will be updated later this decade to GRAVITY+, which will also be installed on ESO’s VLTI and will push the sensitivity further to reveal fainter stars even closer to the black hole. The team aims to eventually find stars so close that their orbits would feel the gravitational effects caused by the black hole’s rotation. ESO’s upcoming Extremely Large Telescope (ELT), under construction in the Chilean Atacama Desert, will further allow the team to measure the velocity of these stars with very high precision. “With GRAVITY+’s and the ELT’s powers combined, we will be able to find out how fast the black hole spins,” says Eisenhauer. “Nobody has been able to do that so far.”


 How to resolve AdBlock issue?
How to resolve AdBlock issue? 



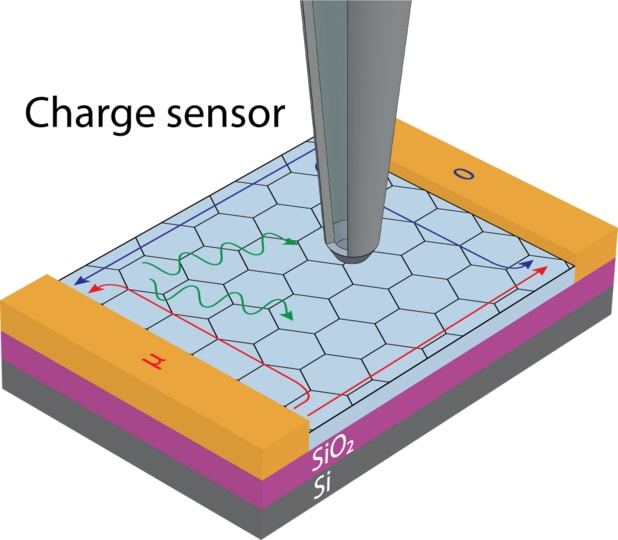 Measuring the properties of a spin-wave is like measuring the properties of a tidal wave if the water itself was undetectable. If you couldn’t see water, how could you measure the speed, height, or the number of tidal waves? One way would be to introduce something into the system that you can measure, like a surfer. The speed of the tidal wave could be detected by measuring the speed of the surfer.
Measuring the properties of a spin-wave is like measuring the properties of a tidal wave if the water itself was undetectable. If you couldn’t see water, how could you measure the speed, height, or the number of tidal waves? One way would be to introduce something into the system that you can measure, like a surfer. The speed of the tidal wave could be detected by measuring the speed of the surfer.