The Gulf Stream system plays a vital role in the climate, and its decline over the past two decades has raised concerns and sparked debates. While the cause of this weakening is uncertain, some simulations suggest that human-induced climate change could be a significant factor in the future. However, a recent study conducted by the GEOMAR Helmholtz Centre for Ocean Research in Kiel, Germany suggests that the observed weakening may be due to natural fluctuations caused by extremely cold winters in the Labrador Sea during the 1990s.
The new supercomputer simulations show that fluctuations in the Labrador Sea can have a significant influence on the strength of sinking processes east of Greenland. An important link is a little-noticed system of deep currents that ensures the rapid spread of Labrador Sea water into the deep-sea basin between Greenland and Iceland. 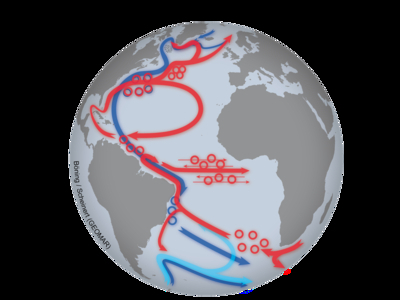
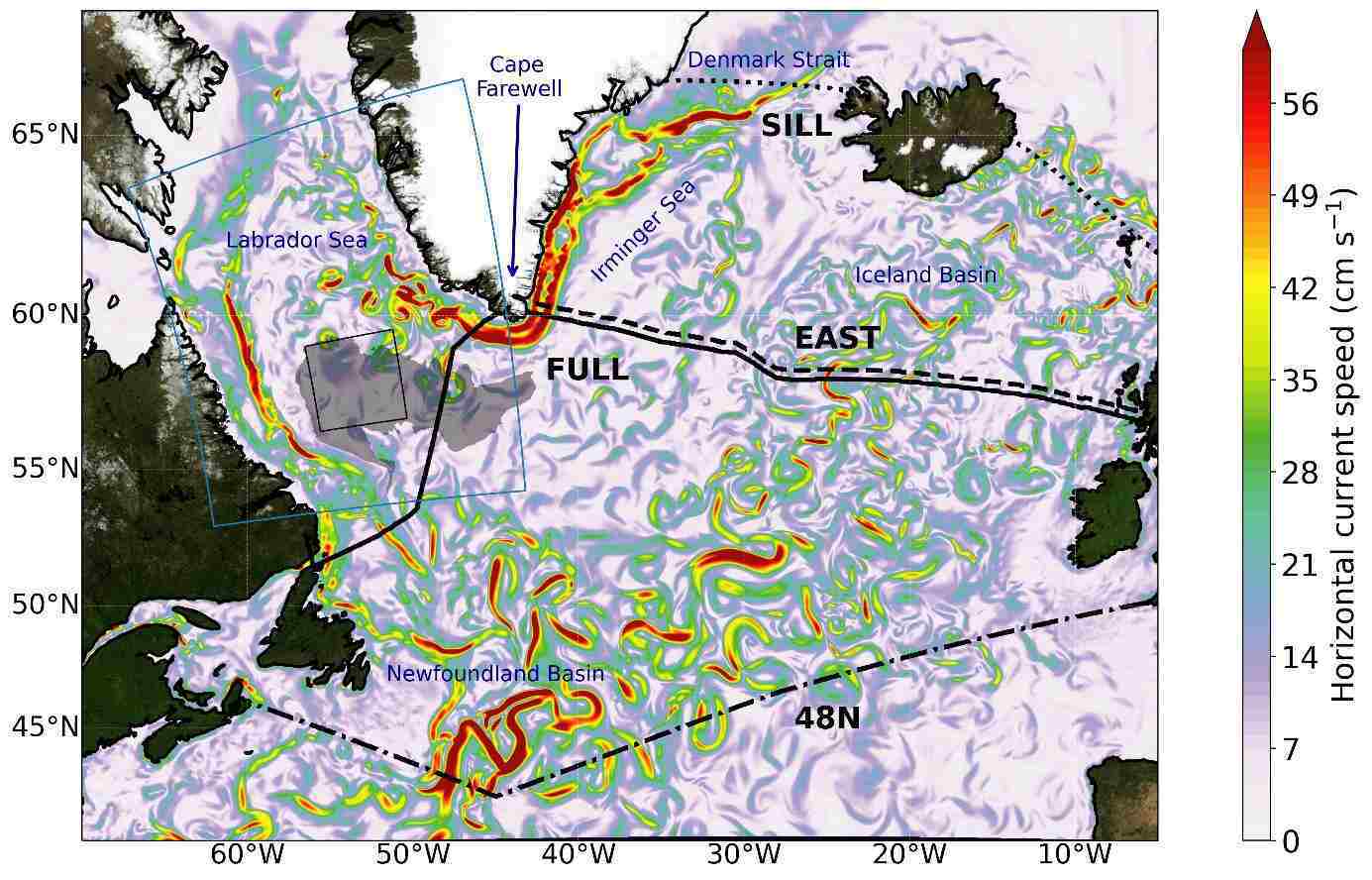
"We oceanographers have long had our eyes on the Labrador Sea between Canada and Greenland," says Professor Dr Claus Böning, who led the study. "Winter storms with icy air cool the ocean temperatures to such an extent that the surface water becomes heavier than the water below. The result is deep winter mixing of the water column, whereby the volume and density of the resulting water mass can vary greatly from year to year."
In the model simulations of the past 60 years, the years 1990 to 1994 have stood out, when the Labrador Sea cooled particularly strongly. "The huge volume of very dense Labrador Sea wateThe Gulf Stream system plays a critical role in the climate, and concerns have been raised about its weakening over the past two decades. While it's unclear whether human-induced climate change has caused these changes, simulations indicate that it's highly probable to occur in the future. However, a recent study by GEOMAR Helmholtz Centre for Ocean Research in Kiel, Germany suggests that the weakening may be due to natural fluctuations caused by extremely cold winters in the Labrador Sea during the 1990s.r that formed following extremely harsh winters led to significantly increased sinking between Greenland and Iceland in the following years," explains Claus Böning. As a result, the model simulations calculated an increase in Atlantic overturning transport of more than 20%, peaking in the late 1990s. The measurements of the circulation in the North Atlantic, which have only been carried out continuously since 2004, would then fall precisely in the decay phase of the simulated transport maximum.
Video: {joomvideos id=326}
"According to our model results, the observed weakening of the Atlantic circulation during this period can therefore be interpreted, at least in part, as an aftereffect of the extreme Labrador Sea winters of the 1990s", summarises Professor Dr. Arne Biastoch, head of the Ocean Dynamics Research Unit at GEOMAR and co-author of the study. However, he clarifies: "Although we cannot yet say whether a longer-term weakening of the overturning is already occurring, all climate models predict a weakening as a result of human-induced climate change as 'very likely' for the future.
Ongoing observing programs and further development of supercomputer simulations are crucial for a better understanding of the key climate-relevant processes. And, of course, for future projections of the Gulf Stream system under climate change.
The research conducted by GEOMAR has revealed a potential link between winter storms over the Labrador Sea and the Gulf Stream system. While the findings are certainly intriguing, further research is needed to confirm the validity of these results. Until then, it is important to remain skeptical.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?