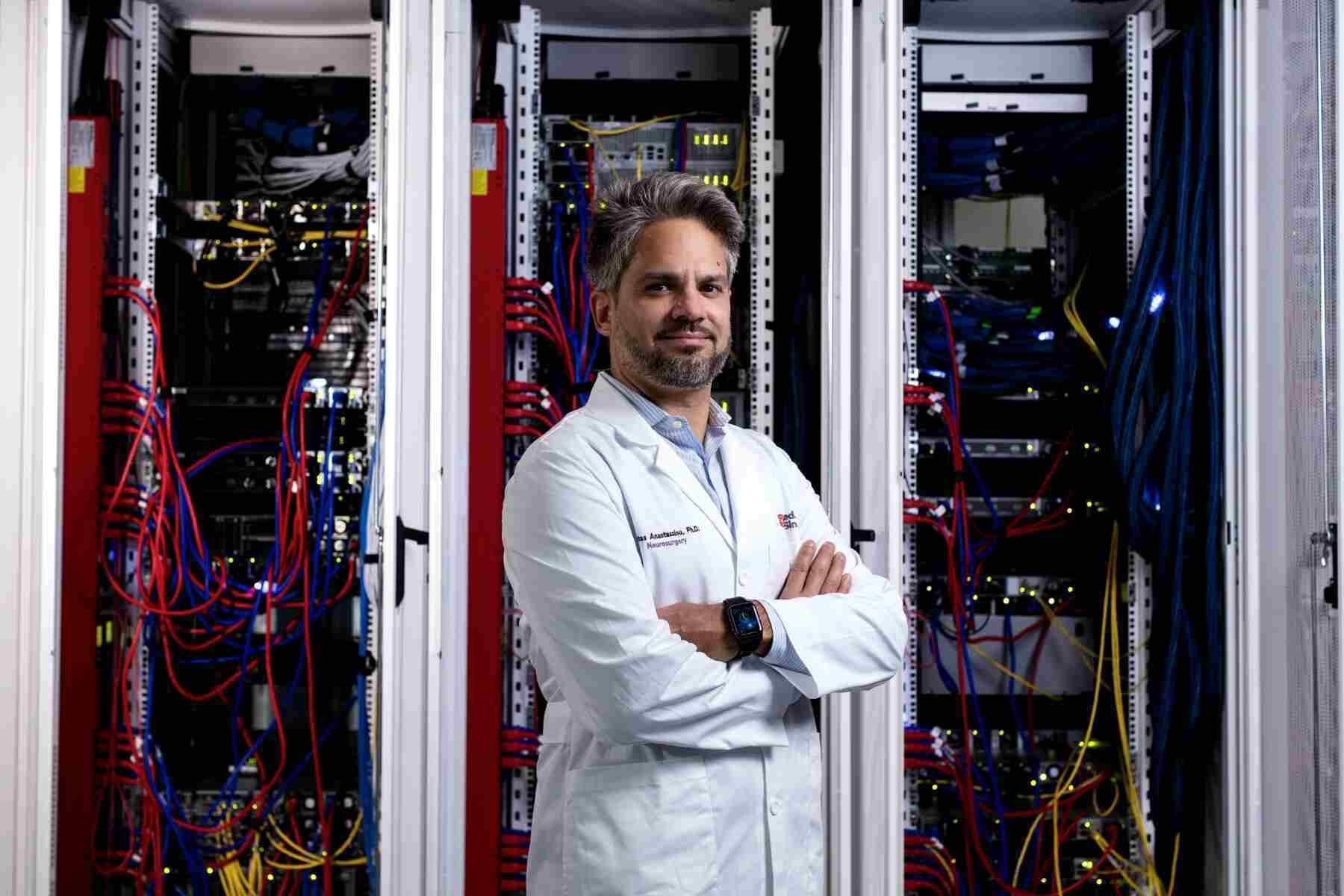
Investigators at Cedars-Sinai have created supercomputer-generated models to bridge the gap between “test tube” data about neurons and the function of those cells in the living brain. Their study could help in the development of treatments for neurological diseases and disorders that target specific neuron types based on their roles. 
“This work allows us to start looking at the brain like the complex machine that it is, rather than as one homogenous piece of tissue,” said Costas Anastassiou, Ph.D., a research scientist in the departments of Neurology, Neurosurgery, and Biomedical Sciences at Cedars-Sinai and senior author of the study. “Once we are able to distinguish between the different cell types, instead of saying that the entire brain has a disease, we can ask which neuron types are affected by the disease and tailor treatments to those neurons.”
Neurons are the main functional units of the brain. The signals passing through these cells—in the form of electrical waves—give rise to all thought, sensation, movement, memory, and emotion.
The study used data from laboratory mice to establish a new method for examining relationships between neuron type and function and focused on the mouse's primary visual cortex, which receives and processes visual information. It is one of the best-studied parts of the brain—both in vitro, where tissue is studied in a dish or test tube outside the living organism, and in vivo, where it is studied in the living animal.
The investigators’ goal was to link the two worlds.
“Based on in vitro studies of genetic makeup and physical structure, we know something about what various classes of neurons look like, but not their function in the living brain,” Anastassiou said. “When we record the activity of brain cells in vivo, we can see what neurons are responding to and what their function is, but not which classes of neurons they are.”
To link form to function, investigators first used in vitro information to create computational models of various types of neurons and to simulate their signaling patterns.
They next took advantage of the newest technology in single-neuron recording to observe activity in the brains of laboratory mice while the mice were exposed to different sorts of visual stimuli. Based on the shapes of the signals or “spikes” of neurons in response to visual input, investigators separated the cells they recorded into six groups.
“Once we had our models and our in vivo data, the fundamental question was which computational models produced the most similar signaling shape and waveform to each of the six in vivo clusters we identified, and vice versa,” Anastassiou said. “Not all of the in vivo clusters and models matched perfectly, but some did.”
More data, and possibly experiments involving more-sophisticated visual stimuli, might be required to match all the computational models and cell clusters, and Anastassiou said that future studies will be dedicated to perfecting the method established in the current paper.
“There’s a wealth of information about the identity of cell types in the human brain, but not about the role of those cell types in cognitive functioning or how they are affected by the disease,” Anastassiou said. “Now there is a window through which we can look at these things and ask these questions. It’s clear that we have a long way to go, but we’re excited about the next steps in this journey.”
The ultimate goal is to pave the way for discoveries that change patients’ lives.
“Our research scientists are continually striving to expand our knowledge of the workings of the human brain at the most detailed level,” said Keith L. Black, MD, chair of the Department of Neurosurgery and the Ruth and Lawrence Harvey Chair in Neuroscience at Cedars-Sinai. “Pinning down the specific type and function of each neuron may one day lead to the discovery of lifesaving treatments for brain diseases and neurological disorders.”
Funding: This research was supported by National Institutes of Health grant number RO1 NS120300-01; National Natural Science Foundation of China grant number 12101570; and Scientific Project of Zhejiang Lab grants 2021KE0PI03, 2022KI0AC01, 2022KI0AC02 and 2022ND0AN01.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?