The alluvial and hydro-climatic changes on the boundary between Huesca and Lleida during the Palaeocene-Eocene global warming are analyzed
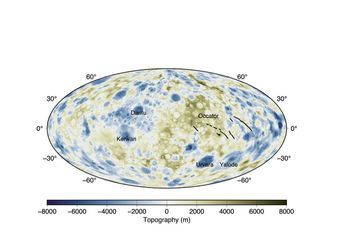
The Department of Geology of the University of the Basque Country UPV/EHU has examined sediments dating back 56 million years in the Tremp-Graus basin (on the border between Lleida and Huesca). It can be deduced from the study that the global warming episode at that time consisted of three phases in which the distribution of precipitation was different. The data from the study can be used to adjust mathematical models used to predict the effects of the current climate change. 
Major carbon emissions into the atmosphere and oceans took place 56 million years ago; that led to intense global warming known as the Palaeocene-Eocene Thermal Maximum and is regarded as an ancient analog of today's anthropogenic warming. “Although the origin or cause of the warming at that time was different, the process was very similar to today’s warming, so it is considered to be similar to today's global warming. The climate is known to have warmed, but other alterations besides warming may occur with climate change. In particular, we wanted to analyze how the hydro-climatic conditions in terms of rainfall changed at that time,” said Aitor Payros, who gained a Ph.D. in Geology at the UPV/EHU.
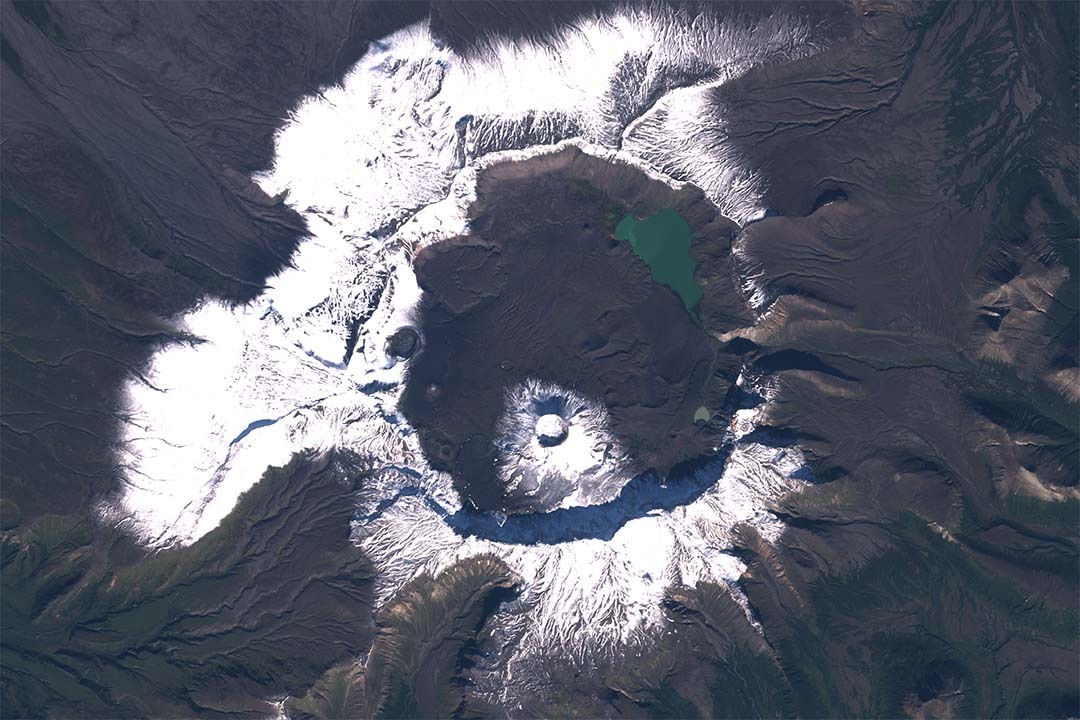
The UPV/EHU’s Department of Geology has investigated the mid-latitude alluvial and hydro-climatic changes recorded in the Tremp-Graus basin (on the border between Lleida and Huesca) during the Palaeocene-Eocene Thermal Maximum and has concluded that what happened then could in some way be similar to what is already happening today in the southeast of the Iberian Peninsula. To do this, they collected historical data from the region and discovered geographical as well as hydro-climatic similarities.
According to Aitor Payros, “we saw that global warming altered the seasonal distribution of precipitation and that it also altered across several phases. At first, rainfall was concentrated in a few months around autumn; later, it became more evenly distributed throughout the year. The last phase, however, tended to be drier. According to Payros, "we can't simply say that global warming is causing temperatures to rise or making rainfall heavier. Things are not that straightforward. Changes do occur, but they are not sustained over the entire period of global warming. Within global warming, there may be several phases".
Looking to the past to predict the future
"We observed that at the onset of that global warming episode there was an increase in seasonal contrasts with respect to precipitation. In other words, precipitation was concentrated around the autumn (with frequent storms and major floods) and during the remaining months, there were periods of drought. And that is precisely what has been happening over the last few decades, and during the last century, in the southeast of the Iberian Peninsula: heavy rainfall is becoming more frequent around autumn and late summer, which was not the case 100 or 200 years ago," said Payros.
The researcher pointed out that it is not possible to predict what will happen in the future in the southeast of the Iberian Peninsula, "but if we assume that the Earth responds in a similar way to the same or similar phenomena, we could surmise that the future annual distribution of precipitation may be more homogeneous in the southeast of the peninsula or other regions with a similar climate".
Payros stresses the potential value of studying palaeo-climates: "We can see what happened millions of years ago. And if what happened is repeated over and over again, in other words, if the Earth always responds in the same way to certain phenomena, we can assume that it will continue to function in the same way in the future.” This type of research can be used to make future predictions: "When the computer or mathematical models used to predict climate are able to reproduce the phenomena that took place during past global warming periods, then they will be able to predict the changes that will occur in the future. Such computer and mathematical models may tally with our data."


 How to resolve AdBlock issue?
How to resolve AdBlock issue?