As the climate warms, wildfires in the Sierra Nevada are happening at unprecedented sizes and intensities, threatening communities and resources throughout Nevada and California. For fire managers trying to understand and predict fire behavior, access to accurate information for decision-making has never been more important.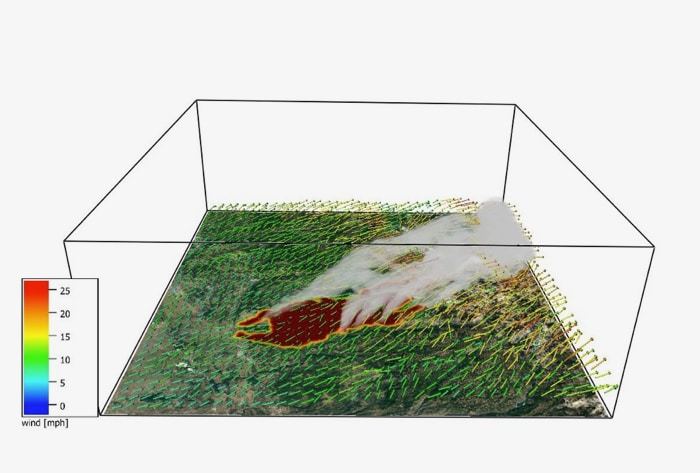 
A generous grant from the NV Energy Foundation will provide $150,000 to support DRI’s development of a Weather and Research Forecast advanced modeling tool that simulates weather, fire, and smoke for firefighting and prescribed fire operations. Forecasts and simulations produced by this model will be available to NV Energy’s fire mitigation team, and other professionals from the prescribed fire and air quality communities in Nevada and California through the work of the California and Nevada Smoke and Air Committee (CANSAC).
“We are committed to protecting our customers and the environment from the increasing risks of natural disasters, which include wildfires,” said Doug Cannon, NV Energy's president and chief executive officer. “The NV Energy Foundation is proud to support DRI in the development of this technology that will help firefighters better assess fire risk and keep our communities safe.”
Funds from the new NV Energy Foundation grant will be used to expand the current high-performance supercomputer system that is used by CANSAC. The system will provide an interface where users such as prescribed fire managers can conduct simulations of fire spread and smoke behavior.
The model will allow for risk assessment of specific locations by modeling different burn scenarios, help meteorologists identify small-scale wind flows that could have adverse effects on fire spread and behavior, and provide critical air quality forecasts for wildfires or burn day decisions. Simulations can be run for near future forecasting (a few days out) or longer-term scenario modeling for projects that might occur a year or more into the future.
“This tool will be useful to wildfire-fighting operations as well as for prescribed fire planning, which is essential to getting some of our fire-adapted ecosystems back into balance,” said Tim Brown, Ph.D., director of DRI’s Western Regional Climate Center. “By supporting the development of this tool, the NV Energy Foundation is providing a great resource to fire managers in Nevada and California and helping to ensure the safety of firefighters and communities across these two states."
“With this generous grant, the NV Energy Foundation will play a key role in developing new technology that will be used to solve real-world problems in fire mitigation and fire safety,” said DRI President Kumud Acharya, Ph.D. “This project is an amazing example of how community organizations like NV Energy can partner with DRI scientists to develop solutions to the problems that face our society and environment.”


 How to resolve AdBlock issue?
How to resolve AdBlock issue?