 Massive neutron stars colliding in space are thought to be able to create precious metals such as gold and platinum. The properties of these stars are still an enigma, but the answer may lie beneath the skin of one of the smallest building blocks on Earth – an atomic nucleus of lead. Getting the nucleus of the atom to reveal the secrets of the strong force that governs the interior of neutron stars has proven difficult. Now a new computer model from the Chalmers University of Technology, Sweden, can provide answers.
Massive neutron stars colliding in space are thought to be able to create precious metals such as gold and platinum. The properties of these stars are still an enigma, but the answer may lie beneath the skin of one of the smallest building blocks on Earth – an atomic nucleus of lead. Getting the nucleus of the atom to reveal the secrets of the strong force that governs the interior of neutron stars has proven difficult. Now a new computer model from the Chalmers University of Technology, Sweden, can provide answers.
The strong force plays the main role
Despite the huge size difference between a microscopic atomic nucleus and a neutron star several kilometers in size, it is largely the same physics that governs their properties. The common denominator is the strong force that holds the particles – the protons and neutrons – together in an atomic nucleus. The same force also prevents a neutron star from collapsing. The strong force is fundamental in the universe, but it is difficult to include in computational models, not least when it comes to heavy neutron-rich atomic nuclei such as lead. Therefore, the researchers have wrestled with many unanswered questions in their challenging calculations.
A reliable way to make calculations
"To understand how the strong force works in the neutron-rich matter, we need meaningful comparisons between theory and experiment. In addition to the observations made in laboratories and with telescopes, reliable theoretical simulations are therefore also needed. Our breakthrough means that we have been able to carry out such calculations for the heaviest stable element – lead,” says Andreas Ekström, Associate Professor at the Department of Physics at Chalmers and one of the main authors of the article. 
The new supercomputer model from Chalmers, developed together with colleagues in North America and England, now shows the way forward. It enables high-precision predictions of properties for the isotope* lead-208 and its so-called ‘neutron skin’.
The thickness of the skin matters
It is the 126 neutrons in the atomic nucleus that form an outer envelope, which can be described as skin. How thick the skin is, is linked to the properties of the strong force. By predicting the thickness of the neutron skin, knowledge can increase about how the strong force works – both in atomic nuclei and in neutron stars.
"We predict that the neutron skin is surprisingly thin, which can provide new insights into the force between the neutrons. A groundbreaking aspect of our model is that it not only provides predictions but also has the ability to assess theoretical margins of error. This is crucial for being able to make scientific progress," says research leader Christian Forssén, Professor at the Department of Physics at Chalmers.
The model used for the spread of the coronavirus
To develop the new computational model, the researchers have combined theories with existing data from experimental studies. The complex calculations have then been combined with a statistical method previously used to simulate the possible spread of the coronavirus.
With the new model for lead, it is now possible to evaluate different assumptions about the strong force. The model also makes it possible to make predictions for other atomic nuclei, from the lightest to the heaviest. 
The breakthrough could lead to much more precise models of, for example, neutron stars and increased knowledge of how these are formed.
"The goal for us is to gain a greater understanding of how the strong force behaves in both neutron stars and atomic nuclei. It takes the research one step closer to understanding how, for example, gold and other elements could be created in neutron stars – and at the end of the day it is about understanding the universe," says Christian Forssén.


 How to resolve AdBlock issue?
How to resolve AdBlock issue? 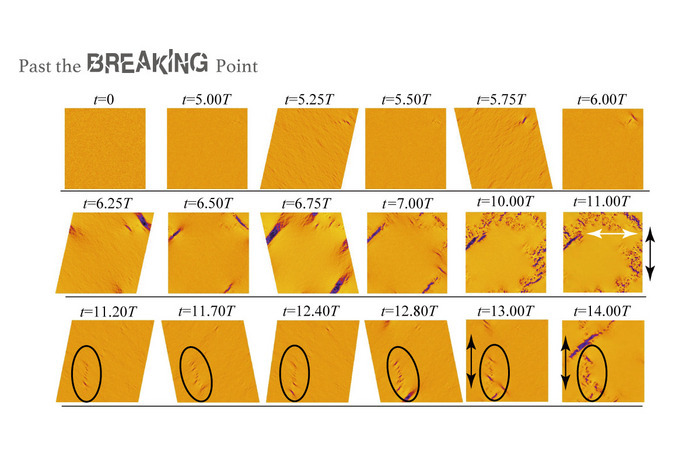 Damage to industrial parts is expensive, results in delays, and may be unsafe to plant workers. But now, scientists from Tokyo Japan have simulated fracture initiated in materials that share a particular physical characteristic and are widely used across domestic, industrial, and scientific applications. Their work showed surprising results that may help prevent damage to industrial parts.
Damage to industrial parts is expensive, results in delays, and may be unsafe to plant workers. But now, scientists from Tokyo Japan have simulated fracture initiated in materials that share a particular physical characteristic and are widely used across domestic, industrial, and scientific applications. Their work showed surprising results that may help prevent damage to industrial parts.