An international team of scientists believes they may have found a molecular mechanism behind the extremely rare blood clots linked to adenovirus COVID-19 vaccines.
Scientists led by a team from Arizona State University, Cardiff University, and others worked with AstraZeneca to investigate vaccine-induced immune thrombotic thrombocytopenia (VITT), also known as thrombosis with thrombocytopenia syndrome (TTS), a life-threatening condition seen in a very small number of people after receiving the Oxford-AstraZeneca or Johnson & Johnson vaccines.
“The mechanism which results in this condition, termed vaccine-induced immune thrombotic thrombocytopenia (VITT), was unknown,” said Abhishek Singharoy, an Arizona State University scientist and corresponding author of the study who teamed up to lead an international effort to tease out the details. So, a team quickly assembled to try to understand the problem more clearly.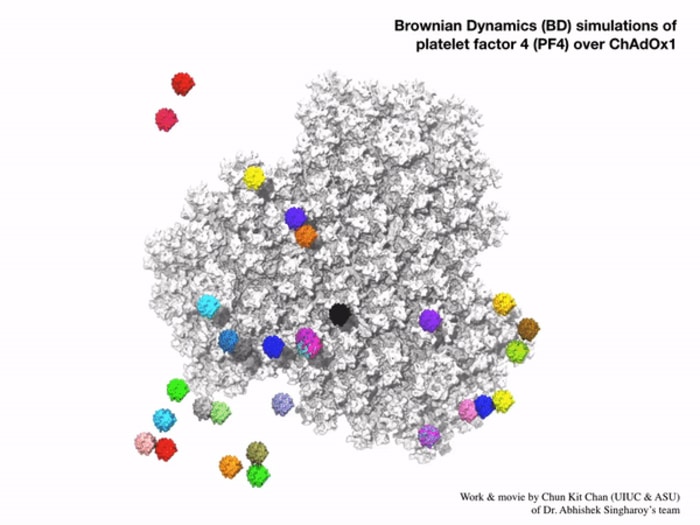 
Together, they worked to solve the structural biology of the vaccine and see the molecular details that may be at play, utilizing ASU’s new cryo-EM facilities, and a state-of-the-art Titan Krios machine at ASU’s Eyring Materials Center at Arizona State University.
ASU scientists included several from the School of Molecular Sciences and Biodesign Institute: Ryan J. Boyd, Daipayan Sarkar, John Vant, Eric Wilson, Chloe D. Truong, Petra Fromme, Po-Lin Chiu, Dewight Williams, and Josh Vermaas (ASU alumnus now at Michigan State University). Mitesh Borad, Bolni M. Nagalo, and Alexander T. Baker were also part of the Arizona-based team.
The global team used state-of-the-art cryo-EM technology to analyze the AstraZeneca vaccine in minute detail to understand whether the ultra-rare side effect could be linked to the viral vector which is used in many vaccines, including those from Oxford/AstraZeneca and Johnson & Johnson.
Their findings suggest it is the viral vector – in this case, an adenovirus used to shuttle the coronavirus’ genetic material into cells – and the way it binds to platelet factor 4 (PF4) once injected that could be the potential mechanism.
In very rare cases, the scientists suggest, the viral vector may enter the bloodstream and bind to PF4, where the immune system then views this complex as foreign. They believe this misplaced immunity could result in the release of antibodies against PF4, which bind to and activate platelets, causing them to cluster together and triggering blood clots in a very small number of people after the vaccine is administered.
“It’s really critical to fully investigate the vector-host interactions of the vaccine at a mechanistic level,” said Singharoy. “This will assist in understanding both how the vaccine generates immunity, and how it may lead to any rare adverse events, such as VITT.”
Their findings are published today in the international journal Science Advances.
Professor Alan Parker, an expert in the use of adenoviruses for medical applications from Cardiff University’s School of Medicine, said: “VITT only happens in extremely rare cases because a chain of complex events needs to take place to trigger this ultra-rare side effect. Our data confirms PF4 can bind to adenoviruses, an important step in unraveling the mechanism underlying VITT. Establishing a mechanism could help to prevent and treat this disorder.”
“We hope our findings can be used to better understand the rare side effects of these new vaccines – and potentially to design new and improved vaccines to turn the tide on this global pandemic.”
Both the AstraZeneca and Johnson & Johnson vaccines use an adenovirus to carry spike proteins from the coronavirus into people to trigger a protective immune response.
When both vaccines showed the ultra-rare side effect of VITT, scientists wondered whether the viral vector had some part to play. Another important clue was that neither the Moderna nor Pfizer vaccines, made from an entirely different technology called mRNA vaccines, showed this effect.
The team used cryo-EM technology to flash-freeze preparations of ChAdOx1, the adenovirus used in the AstraZeneca vaccine, and bombard them with electrons to produce microscopic images of the vaccine components.
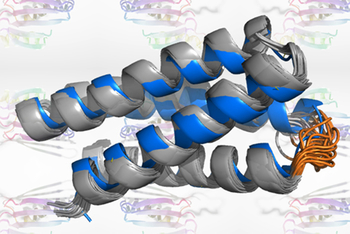
They were then able to look at the atomic level at the structure of the outer protein cage of the virus – the viral capsid – and other critical proteins that allow entry of the virus into the cell.
In particular, the team outlined the details for the structure and receptor of ChAdOx1, which is adapted from chimpanzee adenovirus Y25 – and how it interacts with PF4. They believe it is this specific interaction – and how it is then presented to the immune system – that could prompt the body’s defenses to view it as foreign and release antibodies against this self-protein.
The research team also used the computational models of Singharoy to show that one of the ways the two molecules tightly bind is via electrostatic interactions. The group showed that ChAdOx1 is mostly electronegative. This makes the protein act like the negative end of a battery terminal and could attract other positively charged molecules to its surface.
The first author of the study Dr. Alexander Baker, said: “We found that ChAdOx1 has a strong negative charge. This means the viral vector can act as a magnet and attract proteins with the opposite, positive charge, like PF4.” Baker is a member of ASU’s Biodesign Center for Applied Structural Discovery and an Honorary Research Fellow at Cardiff University School of Medicine.
“We then found that PF4 is just the right size and shape that when it gets close to ChAdOx1 it could bind in between the negatively charged parts of ChAdOx1’s surface, called hexons.”
The research team is hopeful that armed with a better understanding of what may be causing rare VITT they can provide further insights into how vaccines and other therapies, which rely on the same technology, might be altered in the development of the next-generation vaccines and therapies.
“With a better understanding of the mechanism by which PF4 and adenoviruses interact there is an opportunity to engineer the shell of the vaccine, the capsid, to prevent this interaction with PF4. Modifying ChAdOx1 to reduce the negative charge may reduce the chance of causing thrombosis with thrombocytopenia syndrome,” said Baker.
The team likens it to the two birds, one stone effect. The key contacts of individual amino acids that are essential to the capsid protein’s proteins interaction with PF4 can be removed or substituted.
“The modification of the ChAdOx1 hexons to reduce their electronegativity may solve two problems simultaneously: reduce the propensity to cause VITT to even lower levels, and reduce the levels of pre-existing immunity, thus helping to maximize the opportunity to induce robust immune responses, said Singharoy.”
Both the UK-based Medicines and Healthcare products Regulatory Agency (MHRA) and Centers for Disease Control and Prevention (CDC) in the U.S. continue to advise that vaccination is the best way to protect people from COVID-19 and the benefits far outweigh the risk of any known side effects.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?