Findings can help guide surveillance and prevent deadly outbreaks
Since its discovery in 1999, Nipah virus has been reported almost yearly in Southeast Asia, with Bangladesh and India being the hardest hit. In a new study, published today in PLoS Neglected Tropical Diseases, scientists used machine learning to identify bat species with the potential to host Nipah virus, with a focus on India - the site of a 2018 outbreak. Four new bat species were flagged as surveillance priorities.
Barbara Han, a disease ecologist at Cary Institute of Ecosystem Studies, is a co-lead author on the paper. She explains, "While there is a growing understanding that bats play a role in the transmission of Nipah virus in Southeast Asia, less is known about which species pose the most risk. Our goal was to help pinpoint additional species with a high likelihood of carrying Nipah, to target surveillance and protect public health."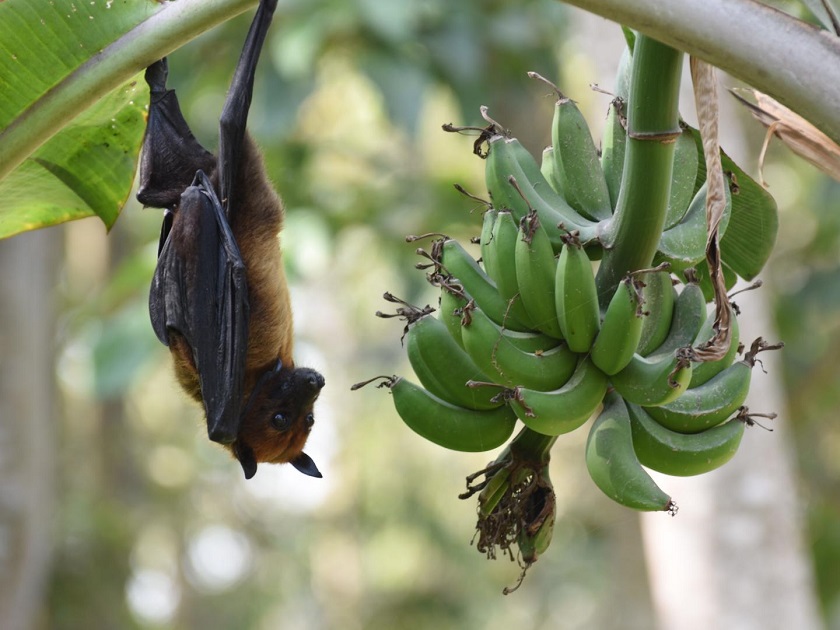  {module In-article}
{module In-article}
Raina K. Plowright, a disease ecologist at Montana State University, was also a co-lead author. She notes, "As this paper was going to press, another case of Nipah virus was confirmed in Kerala. The public health community has again been forced into reactive mode. Our study is a starting point for the research needed to contain Nipah at its source, so we are managing spillover risk, instead of human suffering."
Nipah virus is a highly lethal, emerging henipavirus that can be transmitted to people from the body fluids of infected bats. Eating fruit or drinking date palm sap that has been contaminated by bats has been flagged as a transmission pathway. Once infected, people can spread the virus directly to other people, sparking an outbreak. Domestic pigs are also bridging hosts that can infect people. There is no vaccine and the virus has a high mortality rate.
"Bat-borne viruses are found all over the world, yet surveillance and sampling efforts have been patchy," says Han. "There are likely many competent Nipah hosts that have not been identified. For this reason, there is a need to devise new methods that take all available data into account to guide sampling efforts in India and in other regions."
India is home to an estimated 113 bat species. Just 31 of these species have been sampled for Nipah virus, with 11 found to have antibodies that signal host potential. Plowright notes, "Given the role bats play in transmitting viruses infectious to people, investment in understanding these animals has been low. The last comprehensive and systematic taxonomic study on the bats in India was conducted more than a century ago." 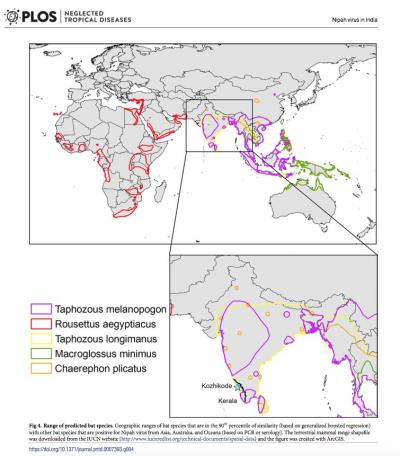 {module In-article}
{module In-article}
Machine learning, a form of artificial intelligence, was used to flag bat species with the potential to harbor Nipah. Han explains, "By looking at the traits of bat species known to carry Nipah globally, our model was able to make predictions about additional bat species residing in India with the potential to carry the virus and transmit it to people. These bats are currently not on the public health radar and are worthy of additional study."
First, the team compiled published data on bat species known to carry Nipah and other henipaviruses globally. Data included 48 traits of 523 bat species, including information on foraging methods, diet, migration behaviors, geographic ranges, and reproduction. They also looked at the environmental conditions in which reported spillovers occurred.
Then they applied a trait-based machine learning approach to a subset of species that occur in Asia, Australia, and Oceana. Their algorithm identified known Nipah-positive bat species with 83% accuracy. It also identified six bat species that occur in Asia, Australia, and Oceana that have traits that could make them competent hosts and should be prioritized for surveillance. Four of these species occur in India, two of which are found in Kerala.
Plowright explains, "We set out to make trait-based predictions of likely henipavirus reservoirs near Kerala. Our focus was narrow, but the model was successful in identifying Nipah hosts, demonstrating that this method could serve as a powerful tool in guiding surveillance for Nipah and other disease systems."
The authors note that their predictions must be combined with local knowledge on bat ecology - including distribution, abundance, and proximity to humans - to design sampling plans that can effectively identify bat hosts that pose a risk to humans. This work provides a list of species to guide early surveillance and should not be taken as a definitive list of reservoirs.
"Surveilling high-risk bat populations can provide early warning for veterinarians and public health authorities to take preventative measures needed to preempt an outbreak. Identifying which species harbor disease is an important first step in surveillance planning. We also need to prioritize research on which virus strains pose the greatest risk to people. Ultimately, the goal is to extinguish risk, not fight fires," Han concludes.
 {module In-article}
{module In-article}

 How to resolve AdBlock issue?
How to resolve AdBlock issue?  {module In-article}
{module In-article} {module In-article}
{module In-article} {module In-article}
{module In-article}