There are no scales for weighing black holes. Yet astrophysicists from the Moscow Institute of Physics and Technology have devised a new way for indirectly measuring the mass of a black hole, while also confirming its existence. They tested the new method, reported in the Monthly Notices of the Royal Astronomical Society, on the Messier 87 active galaxy.
Active galactic nuclei are among the brightest and most mysterious objects in space. A galaxy is deemed active if it produces a thin long beam of matter and energy directed outward. Known as a relativistic jet, this phenomenon cannot be accounted for by the stars in the galaxy. The current consensus is that the jets are produced by some kind of "motors," termed galactic nuclei. While their nature is poorly understood, researchers believe that a spinning black hole could power an active galaxy.
Messier 87 in the Virgo constellation is an active galaxy that is closest to Earth, and also the one best studied. It has been observed on a regular basis since 1781, when it was first discovered as a nebula. It took some time before astronomers realized that it was a galaxy, and its optical jet -- discovered in 1918 -- was the first one ever to be observed.
The structure of the Messier 87 jet has been meticulously studied, with its plasma jet velocities mapped and the temperature and particle number density near the jet measured. The jet's boundary has been studied in such fine detail that researchers discovered it was inhomogeneous along its length, changing its shape from parabolic to conical. Originally discovered as an isolated case, this effect was later confirmed for a dozen other galaxies, though M87 remains the clearest example of the phenomenon.
The sheer bulk of observations allow for testing hypotheses regarding the structure of active galaxies, including the relation between the jet shape break and the black hole's gravitational influence. Jet behavior and the existence of the supermassive black hole are two sides of the same coin: The former can be explained in terms of the latter while theoretical models of black holes are tested via jet observations.
Astrophysicists exploited the fact that the jet boundary is made up of segments of two distinct curves and used the distance between the core and the break of the jet, together with the jet's width, to indirectly measure the black hole mass and spin. To that end, MIPT scientists developed a method that combines a theoretical model, supercomputer calculations, and telescope observations. 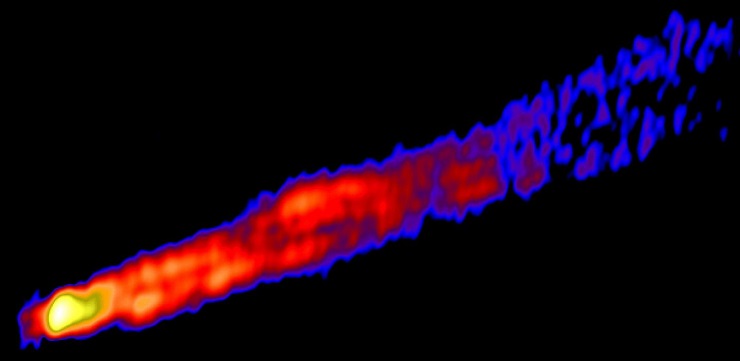 {module In-article}
{module In-article}
The researchers are trying to describe the jet as a flow of magnetized fluid. In this case, the shape of the jet is determined by the electromagnetic field in it, which in turn depends on various factors, such as the speed and charge of jet particles, the electric current within the jet, and the rate at which the black hole accretes matter. A complex interplay between these characteristics and physical phenomena gives rise to the observed break.
There is a theoretical model that predicts the break, so the team could determine which black hole mass results in the model reproducing the observed shape of the jet. This provided a new model for black hole mass estimation, a new measurement method, and a confirmation of the hypotheses underlying the theoretical model.
"The new independent method for estimation of black hole mass and spin is the key result of our work. Even though its accuracy is comparable to that of the existing methods, it has an advantage in that it brings us closer to the end goal. Namely, refining the parameters of the core 'motor' to deeper understand its nature," said Elena Nokhrina, the lead author of the paper and deputy head of the MIPT laboratory involved in the study.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?  {module In-article}
{module In-article} {module In-article}
{module In-article}