Methicillin-resistant Staphylococcus aureus (MRSA) is a bacterial infection that has become resistant to most of the antibiotics used to treat regular staph infections. 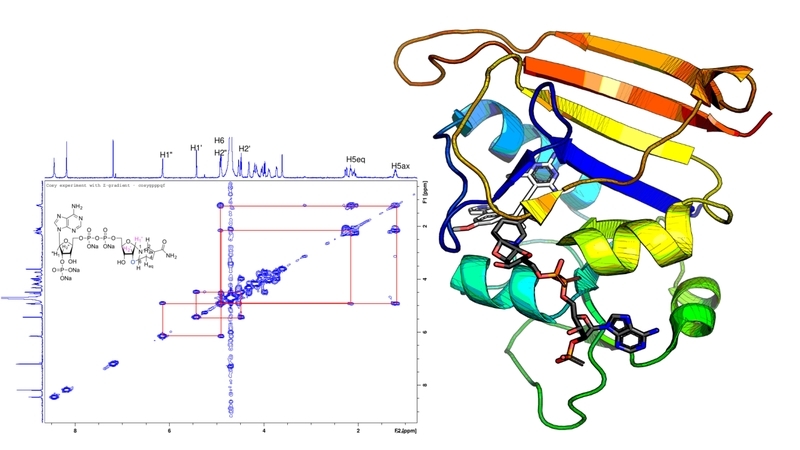
Bruce Donald, Ph.D., James B Duke Distinguished Professor of Computer Science, and collaborators at the University of Connecticut are working to develop new enzyme inhibitors to fight MRSA. In research published in PLOS Computational Biology, the team discovered how a single small mutation makes a big difference in drug efficacy.
They examined dihydrofolate reductase (DHFR), an enzyme that antibiotics target to fight MRSA. Drugs that inhibit DHFR work a bit like locks and keys; they bind to enzymes in MRSA, which have a specific three-dimensional structure that only allows molecules that fit precisely to attach to them.
A mutation can change the structure of a bacterial enzyme and cause drugs to lose effectiveness. The F98Y mutation is a well-known resistance mutation. A slight change in the 98th amino acid in the DHFR enzyme changes phenylalanine to tyrosine. “Those two amino acids are structurally similar,” said Graham Holt, grad student in the Donald lab, “but the mutation has a huge effect on the efficacy of the inhibitors.” In essence, it changes the lock.
Pablo Gainza, Ph.D., a former graduate student in the Donald lab, thought he should be able to predict this mutation using OSPREY, a suite of programs for computational structure-based protein design developed in the Donald lab. But he couldn’t. After knocking down hypothesis after hypothesis to figure out why he was unable to predict this mutation, he went back to examine the starting structure.
“We looked at the electron density data from the crystallographer and found something strange,” Donald said. In trying to determine the structure of the F98Y mutant, crystallographers used a computer program that—unbeknownst to them – flipped the chirality, or made a mirror image, of the NADPH cofactor to get a better fit. The “flipped” chemical species they discovered through their analysis exists in experimental conditions in the laboratory and plausibly in vivo.
“Using OSPREY, we discovered this flipped chirality,” Donald said, “which we believe happened because of the F98Y mutation.” As in 2-factor authentication, the single enzyme mutation and the flipped cofactor appear to conspire together to evade the inhibitor.
This “chiral evasion” changes the structural basis for resistance. But now Donald and colleagues know not only how a single small mutation changed the lock, but also the structure they need to make a better key – a better drug inhibitor.
“This is the first example of an enzyme that exploits the chirality of its cofactor to evade its inhibitors,” Holt said. “Now that we see this happening, that will help inform computational strategies to develop better inhibitors.”
The Donald lab showed that, by taking flipped chirality into account, OSPREY’s predictions closely match experimental measurements of inhibitor potency. They worked with collaborators at the University of Connecticut who conducted biochemical experiments to test the theory and provide structural evidence.
“This is only the beginning of the story,” Donald said. “Our discovery of chiral evasion should lead to more resilient inhibitors: better drug designs.” Right now, most drug design is reactive, waiting for resistance to arise, which it always does. “We hope to make drug design proactive, by using our algorithms to anticipate resistance,” Donald said.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?