NASA’s Near Space Network enables spacecraft exploring the solar system and Earth to send back essential science data for researchers and scientists to investigate and make profound discoveries.
Now, the network has integrated four new global antennas to further support science and exploration missions. In December 2022, antennas in Fairbanks, Alaska; Wallops Island, Virginia; Punta Arenas, Chile; and Svalbard, Norway went online to provide present and future missions with S-, X-, and Ka-band communications capabilities. 
These new antennas were created to support missions capturing immense amounts of data. Just as scientists increase their instrument capabilities, NASA also advances its communications systems to enable missions near Earth and in deep space.
This upgrade is bringing unprecedented flexibility to the Near Space Network and will enhance direct-to-Earth communications – the process by which a satellite takes a picture and then sends the image over radio waves to an antenna on Earth. This data is then processed and sent to scientists. The Near Space Network is managed by NASA’s Space Communications and Navigation (SCaN) program office, which oversees the development and enhancement of NASA’s two primary communications networks: the Near Space and Deep Space networks.
The Near Space Network provides missions with communications services through a blend of government-owned and commercial assets. To develop these new antennas, the team worked with commercial partner Kongsberg Satellite Services (KSAT), who created the Chile and Norway antennas, while NASA developed the other two in Virginia and Alaska.
Now operational, the four antennas are integrated into the network’s service catalog, advancing its capabilities to support science and exploration missions that use enhanced instrumentation. Now, missions using the network will be able to send back terabytes of data for processing and discovery.
An example is the upcoming Plankton, Aerosol, Clouds, ocean Ecosystem (PACE) mission, which will help researchers better understand ocean ecosystems and carbon cycling and reveal how aerosols might fuel phytoplankton growth on the ocean’s surface.
“Missions like the PACE satellite incorporate high-resolution science instruments,” said Damaris Guevara, project lead for the networking upgrade. “These instruments require advanced space communications capabilities, like Ka-band, to get the entirety of their data back to Earth.”
The new antennas also will have new networking capabilities.
All four ground stations are incorporating Delay/Disruption Tolerant Networking (DTN). DTN will empower missions with unparalleled connectivity by storing and forwarding data at points along the network to ensure critical information reaches its destination. DTN is an advanced communications capability being developed and tested by NASA’s SCaN and Space Technology Mission Directorate.
Additionally, to enhance mission teams’ access to data, the network incorporates cloud-based data storage services. Satellites like PACE will downlink their data to an antenna, and that data will go through the ground station’s high-rate data processors to a cloud-based storage and data access service that will allow mission teams to acquire their data faster and from almost anywhere. This reduces hardware needs and lowers overall storage costs.
Multiple missions will benefit from this new infrastructure and advanced capabilities, including the NASA-Indian Space Research Organization Synthetic Aperture Radar (NISAR) satellite. Launching in 2024, NISAR will measure Earth’s changing ecosystems, dynamic surfaces, ice masses, and more.
With four new antennas around the globe, the Near Space Network is advancing its capabilities to support science and exploration missions that use enhanced instrumentation. Now, missions using the network will be able to send back terabytes of data for processing and discovery.


 How to resolve AdBlock issue?
How to resolve AdBlock issue? 
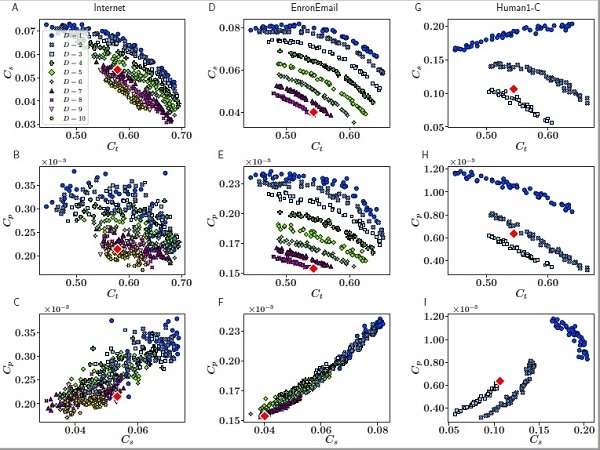 Reducing redundant information to find simplifying patterns in data sets and complex networks is a scientific challenge in many knowledge fields. Moreover, detecting the dimensionality of the data is still a hard-to-solve problem. A new study presents a method to infer the dimensionality of complex networks through the application of hyperbolic geometrics, which capture the complexity of relational structures of the real world in many diverse domains.
Reducing redundant information to find simplifying patterns in data sets and complex networks is a scientific challenge in many knowledge fields. Moreover, detecting the dimensionality of the data is still a hard-to-solve problem. A new study presents a method to infer the dimensionality of complex networks through the application of hyperbolic geometrics, which capture the complexity of relational structures of the real world in many diverse domains.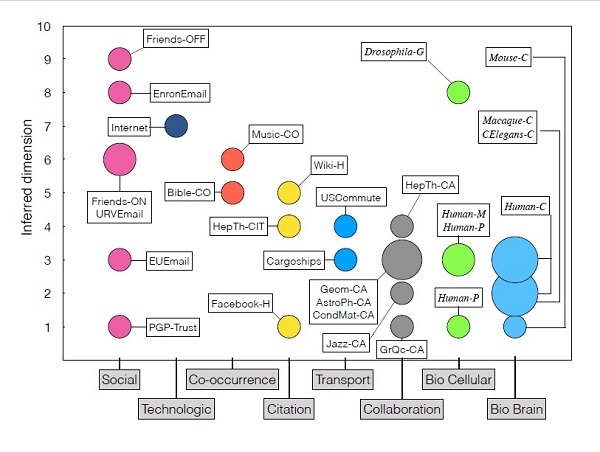 What is the real dimensionality of social networks and the Internet?
What is the real dimensionality of social networks and the Internet?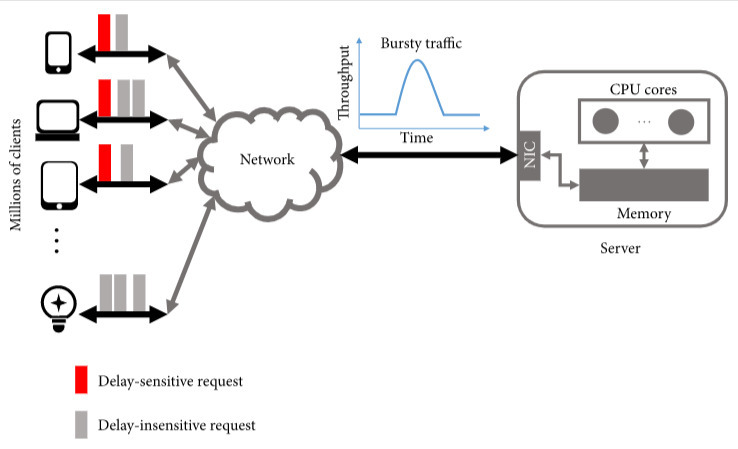 Network interaction has become ubiquitous with the development of the information age and has penetrated into every field of our lives, such as cloud gaming, web searching, and autonomous driving, promoting human progress and providing convenience to society. However, the growing number of clients also caused some problems affecting the user experience. Online services may not respond to some users within the expected timeframe, which is known as high tail latency. In addition, the server's burst traffic exacerbates this issue.
Network interaction has become ubiquitous with the development of the information age and has penetrated into every field of our lives, such as cloud gaming, web searching, and autonomous driving, promoting human progress and providing convenience to society. However, the growing number of clients also caused some problems affecting the user experience. Online services may not respond to some users within the expected timeframe, which is known as high tail latency. In addition, the server's burst traffic exacerbates this issue.