A Kobe University research group has shed light on how low-frequency tectonic tremors occur; these findings will contribute to better predictions of future megathrust earthquakes. 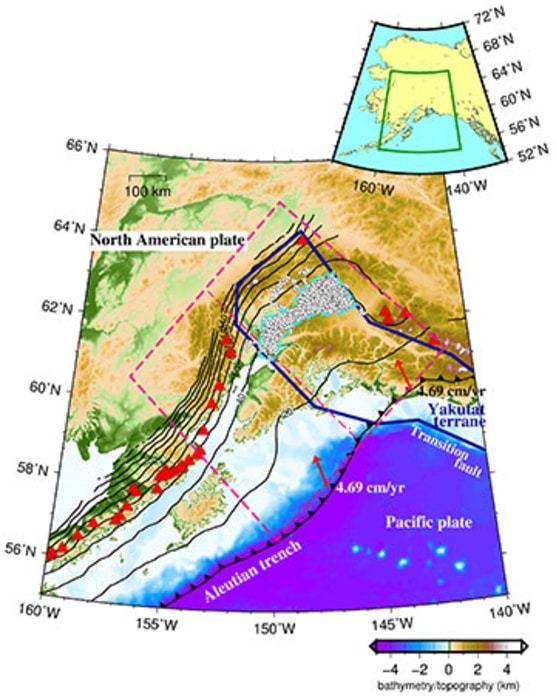
In addition to the subducting Pacific plate, the Alaska subduction zone is also characterized by a subducting oceanic plateau called the Yakutat terrane. Low-frequency tectonic tremors, which are a type of slow earthquake, have only been detected in the subducted Yakutat terrane area. However, the mechanism by which these events occur is not well understood.
Researchers at Kobe University performed a 3D numerical thermomechanical simulation of thermal convection in the Alaska subduction zone to reveal the mechanism behind these low-frequency tremors. Based on the 3D thermal structure obtained from the simulation, and the indications of hydrous minerals contained in the slab, the researchers calculated the water content distribution and compared the results of these calculations in the area where the tremors occur.
The results revealed high levels of dehydration in the marine sediment layers and ocean crust in the earthquake region. The researchers believe that the reason the tremors only occur in the Yakutat terrane is that the marine sediment layers and ocean crust are thicker there, which means that the level of dehydration is higher than in the western adjacent Pacific plate (where tectonic tremors don’t have to occur).
The Kobe University research group consisted of 2nd year Master’s student IWAMOTO Kaya (Department of Planetology, Graduate School of Science), Academic Researcher SUENAGA Nobuaki, and Professor YOSHIDA Shoichi (both of the Research Center's for Urban Safety and Security).
Main Points
- Elucidating the mechanism by which low-frequency tremors occur is important for understanding the plate subduction process. It is believed that this will also help illuminate how shallower megathrust earthquakes occur.
- In this study, the research group constructed a 3D thermomechanical model of the Alaska subduction zone and calculated the subducting plate’s maximum water content and level of dehydration.
- The dehydration levels from the subducting plate’s marine sediment layers and ocean crust were highest in the region where low-frequency tremors occur. Therefore, it is thought that the water expelled from the subducted plate contributes to the occurrence of these tectonic tremors.
Research Background
An oceanic plateau called the Yakutat terrane is subducting in the Alaska subduction zone. Low-frequency tectonic tremors occur at this subducting plateau. The region where slow earthquakes (such as low-frequency tectonic tremors) occur is deeper and adjacent to the area where megathrust earthquakes occur, which suggests a connection between the two. Revealing the mechanism behind how low-frequency tectonic tremors occur is therefore important for understanding the occurrence of various earthquake events in subduction zones. This research group constructed a 3D thermomechanical model of the Alaska subduction zone so that they could investigate the temperature and level of dehydration in the areas near where low-frequency tremors occur.
Research Methodology
The researchers performed a 3D numerical thermomechanical simulation by the subduction of the Yakutat terrane and Pacific plate in the Alaska subduction zone. It is thought that as the Pacific plate subducts, it brings the hydrous minerals in the slab into the deep high temperature and high-pressure regions, and these conditions cause a dehydration reaction where water is expelled from the hydrous minerals. Based on the 3D thermal structure obtained from the numerical simulation, the researchers determined the dehydration levels of the hydrous minerals in the slab. From these results, it was understood that in the region where low-frequency tremors occur, a large amount of water is expelled due to the high temperature and high-pressure conditions that cause the dehydration degradation reactions. It is thought that low-frequency earthquakes don’t occur in the Pacific plate because it has thin layers and therefore experiences little dehydration. On the other hand, the Yakutat terrane’s ocean crust and marine sediment layers are comparatively thicker, meaning that it experiences high levels of dehydration. The researchers concluded that this is why low-frequency tectonic tremors only occur in the Yakutat terrane.
Further Research
In 1964, a megathrust earthquake occurred in Alaska. This is the biggest earthquake that has occurred in the Alaska subduction zone and the second most powerful earthquake recorded in world history. The low-frequency tectonic tremors that were the subject of this research occurred close to the epicenter of the 1964 earthquake, at the downdip of the plate interface.
Next, the research group will continue to make thermomechanical models of various subduction zones search for universal and regional characteristics of the causal mechanisms behind undersea megathrust earthquakes and slow earthquakes. This research will contribute to improving our understanding of how earthquakes occur and our ability to predict future megathrust earthquakes.


 How to resolve AdBlock issue?
How to resolve AdBlock issue? 
