Withered meadows and fields, dry stream beds, dead forests, and reduced power plant outputs – the drought years of 2018, 2019, and 2020 were exceptional and had substantial impacts on nature and the economy. Previously it was not clear where they should be classified in their historical dimension. Now we know: "The 2018 to 2020 drought sets a new benchmark for droughts in Europe", says Dr. Oldrich Rakovec, UFZ modeler and lead author of the article published in the Earth’s Future journal of the American Geophysical Union. The scientists documented this with an extensive compilation of data and modeling techniques which allowed them to reconstruct historical droughts back to 1766 and compare their extents with the drought of 2018 to 2020.
The drought from 2018 to 2020 thus affected approximately one-third of the land area of Europe, especially in central Europe, such as Germany, France, and the Czech Republic. "No other drought event over the last 250 years had such a large spatial extent as this one", explains Oldrich Rakovec. The total duration of the drought event in Europe was also unusually long, starting in April 2018 and not ending until December 2020: 33 months. Only the drought between 1857 and 1860 lasted slightly longer for a total of 35 months. What’s more: The drought from 2018 to 2020 also continued in 2021 and 2022 in deeper soils (i.e., up to 2m below the ground surface). "Although 2021 was wetter and supplied much-needed water in the upper soil important for sustainable agriculture activities, the moisture did not penetrate to greater depths", says the UFZ modeler.
The average drought duration was also unusually long in the 50 x 50 km grid cells in which the scientists subdivided Europe for their modeling activity. Because a drought event develops dynamically in space and time (i.e., it starts at one point, then continues developing, and finally ends somewhere else) its mean duration differs from its total one. In this case, the 2018-2020 event exhibited a mean drought duration of 12 months. 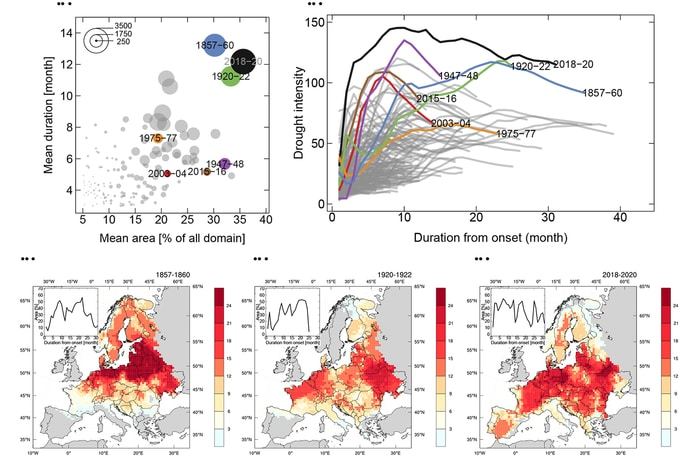
In the past, only the drought event from 1857 to 1860 lasted longer, with a mean duration of 13 months. The scientists define drought as the time in which the current soil-water content in the top 2-m soil falls below the level that has been reached only 20 percent of the time during the 250 years. To reconstruct these historical droughts, the scientists used the mHM hydrologic model developed at the UFZ. Among other things, this environmental model can be used to estimate soil moisture content based on past temperature and precipitation records.
The rise in air temperature also reached a historical record during the 2018-2020 drought event, with an anomaly of 2.8 degrees Celsius above the long-term average over the past 250 years. "The droughts in the past were colder than recent droughts in which the average temperature hardly changed", says Dr. Rohini Kumar, UFZ modeler and co-author of the article. The effects of a drought event become significantly more severe if, in addition to the precipitation deficit (approximately 20 percent for major drought events in past centuries), the warmer conditions prevail. This combined effect results in greater evaporation losses, leading to declining soil-water levels. The scientists also examined the consequences of the lack of water for agriculture during this drought event. They compared average annual crop yields for wheat, grain maize, and barley, between 2018 and 2020 with those between 1961 and 2021. The results indicate that harvests were significantly reduced in countries affected primarily by the 2018-2020 drought. For example, grain maize production decreased between 20 and 40 percent in the Benelux countries, Germany, and France; wheat reduced by up to 17.5 percent in Germany, and barley reduced by 10 percent in nearly all of Europe.
How droughts will develop in Europe in the future also depends on the severity of global warming. The scientists modeled the potential extent and duration of droughts for two representative concentration pathways (RCPs), which describe whether the future greenhouse gas emission scenarios will be more moderate (RCP4.5) or will continue unhindered (RCP8.5) up to the year 2100. The scientists determined that the mean drought duration increases significantly to up to 100 months for an RCP4.5 scenario, while the drought areal extent is projected to increase, covering up to 50 percent of Europe. The situation is different for the extreme RCP8.5 scenario: In this case, the mean drought duration could be more than 200 months, and the areal extent could affect up to 70 percent of Europe. "Decision-makers should be prepared for significantly more severe drought events in the future. Especially for devising new agricultural policies, this should be considered as a wake-up call to assess suitable measures to mitigate the threatening lack of water", says Dr. Luis Samaniego, co-author of the article and Head of the Stochastic and Land Surface Hydrology Working Group at the UFZ. On a regional basis, this could be the establishment of large water reservoirs, such as underground storage systems; intelligent and smart irrigation technologies, or the breeding of more heat-resistant cultivars.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?