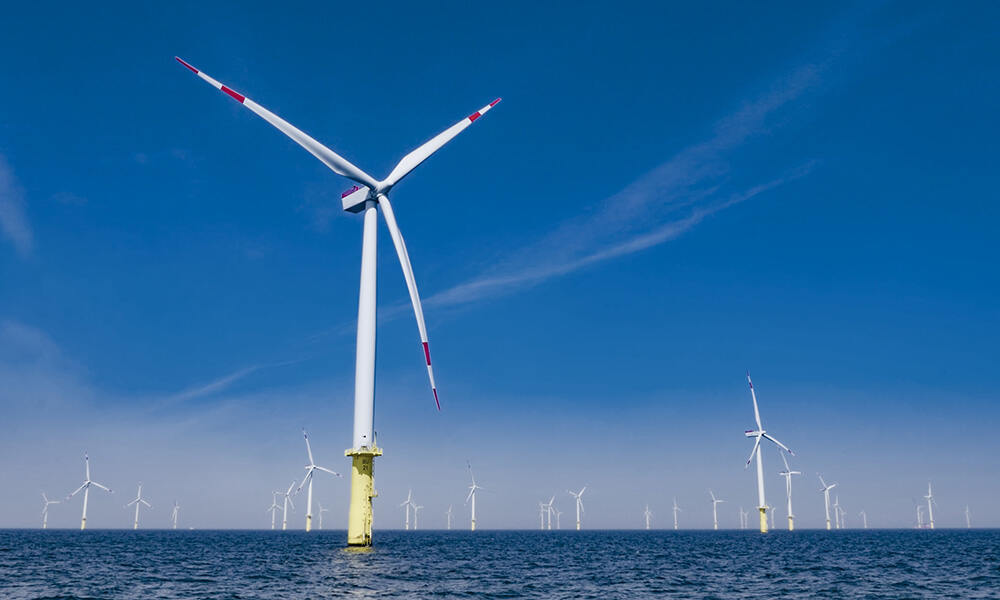
Wind farms have been hailed as a promising source of renewable energy. However, new research by the University of British Columbia Okanagan (UBCO) and Delft University of Technology (TU Delft) in the Netherlands has raised concerns about their effectiveness. The researchers used supercomputer simulations to study the impact of wind farms on air patterns. Their findings have implications for wind farm productivity and the environment.
The researchers developed a modeling framework called the Toolbox for Stratified Convective Atmospheres (TOSCA) to study how wind farms affect the movement of air. They aimed to improve wind energy forecasts and increase productivity. However, when they examined how large wind farms impact natural wind patterns, they found that the results were not as positive as expected.
Dr. Joshua Brinkerhoff, an Associate Professor in UBCO's School of Engineering, explains that wind farms can alter the structure of incoming wind. This structure, known as the atmospheric boundary layer, monitors the wind's speed, temperature, and pressure at different altitudes. The researchers argue that wind farms' alteration of this layer has significant implications for their power output.
Dr. Brinkerhoff emphasizes the importance of proper wind farm design. Poorly designed wind farms can generate less power than expected, making them economically unviable. While software assists in the placement of turbines to maximize output, the researchers argue that their modeling framework is a valuable tool for engineers to design more effective wind farms.
However, skeptics argue that computer modeling may not capture the complex interactions between wind farms and the environment accurately. The lack of precision in estimating power production has significant financial repercussions for wind farm operators. The overestimation of energy output, a common issue not adequately captured by current models, becomes financially disastrous.
The research team acknowledges that their modeling framework, TOSCA, can help forecast the efficiency of wind farms during their establishment. Yet, critics contend that relying solely on simulated data to determine power estimates may not provide an accurate representation of real-world conditions. Skepticism remains regarding the translation of simulation results into practical outcomes.
Although supercomputer simulations represent an advancement in our understanding of wind farm dynamics, it is crucial to consider diverse perspectives on their effectiveness. The interaction between wind farms and the atmosphere is a complex phenomenon that requires a multidisciplinary approach, combining computational models with empirical studies and real-world data.
This research was supported by Mitacs Globalink, UL Renewables, and the Natural Science and Engineering Research Council of Canada. This research aimed to address the challenges facing wind energy. Computational resources were provided by the Digital Research Alliance of Canada and Advanced Research Computing at the University of British Columbia.
As the debate surrounding wind farms and their impact on the environment and energy production continues, it is clear that further research and a holistic understanding of these complex systems are required. Only through careful consideration of the limitations and uncertainties of supercomputer simulations can we arrive at truly sustainable solutions for our energy needs.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?