A new study conducted by Hebrew University provides ground-breaking insights into Tidal Disruption Events
The enigmatic nature of supermassive black holes has fascinated astronomers for years, as they offer a glimpse into the depths of our universe. A recent study conducted by Dr. Elad Steinberg and Dr. Nicholas C. Stone at the Racah Institute of Physics, The Hebrew University, Jerusalem, Israel, has revealed new insights into these cosmic giants through the use of supercomputer simulations.
Supermassive black holes, which can weigh millions to billions of times that of our Sun, remain incomprehensible, despite their central role in shaping galaxies. The immense gravity they generate warps the fabric of spacetime, creating an environment that defies our understanding, and presents a challenge for observational astronomers.
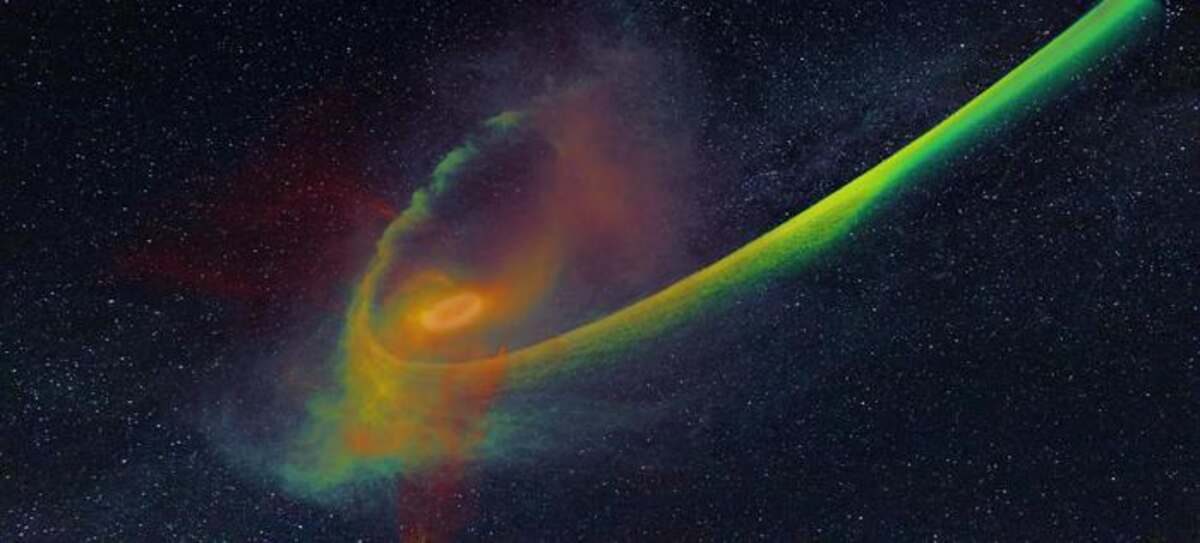
Tidal Disruption Events (TDEs) are dramatic phenomena that occur when unlucky stars get too close to a black hole's event horizon, only to be ripped apart into thin streams of plasma. As this plasma returns towards the black hole, a series of shockwaves heat it, which results in an extraordinary display of luminosity—a flare that exceeds the collective brightness of an entire galaxy for weeks or even months.
Dr. Steinberg and Dr. Stone's study represents a significant advancement in unraveling the mysteries of these cosmic events. Their groundbreaking work meticulously recreates a realistic TDE, capturing the entire sequence from the initial disruption of the star to the pinnacle of the ensuing luminous flare. This achievement is made possible by pioneering radiation-hydrodynamics simulation software developed by Dr. Steinberg at The Hebrew University.
Their research has revealed an unexplored type of shockwave within TDEs, which dissipates energy at a faster rate than previously thought. By shedding light on this aspect, the study resolves a longstanding theoretical debate and confirms that the brightest phases of a TDE flare are powered by shock dissipation.
The implications of these findings are profound. They pave the way for precise measurements of crucial black hole properties, such as mass and spin, and serve as a litmus test for validating Einstein's predictions in extreme gravitational environments. TDE observations hold tremendous potential, enabling scientists to decode the fundamental workings of supermassive black holes and unlock the celestial mysteries that lie at the heart of galaxies.
This remarkable research highlights the transformative power of supercomputer simulations in deciphering the secrets of the universe. Dr. Steinberg and Dr. Stone's simulations represent a significant milestone in our quest to unravel the intricate dynamics of TDEs and comprehend the fundamental workings of supermassive black holes.
As we delve deeper into the mysteries of the cosmos, it is crucial to embrace diverse perspectives and collaborative efforts. The Hebrew University's study exemplifies the power of teamwork and innovation in unraveling the complexities of our universe. By leveraging the computational capabilities of supercomputers, scientists from different backgrounds can synergize their expertise, revolutionizing our understanding of the cosmos.
As we celebrate this remarkable achievement, let us be inspired by the vast possibilities that lie ahead. The journey of exploration continues, with supercomputer simulations serving as our guiding light, illuminating the darkest corners of the cosmos and kindling the sparks of inspiration for future generations of astronomers and researchers.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?