In the vast universe, there are realms beyond our imagination, and NASA's Hubble Space Telescope has once again brought us one step closer to understanding these cosmic wonders. Recent observations from Hubble have revealed the awe-inspiring transformation of an exoplanet's atmosphere over the course of three years. This groundbreaking discovery not only sheds light on the dynamic nature of distant worlds but also brings us closer to identifying potentially habitable exoplanets with stable climates. Let us embark on a journey through the lens of Hubble to unravel the mysteries of the cosmos.
Witnessing the Dance of Nature:
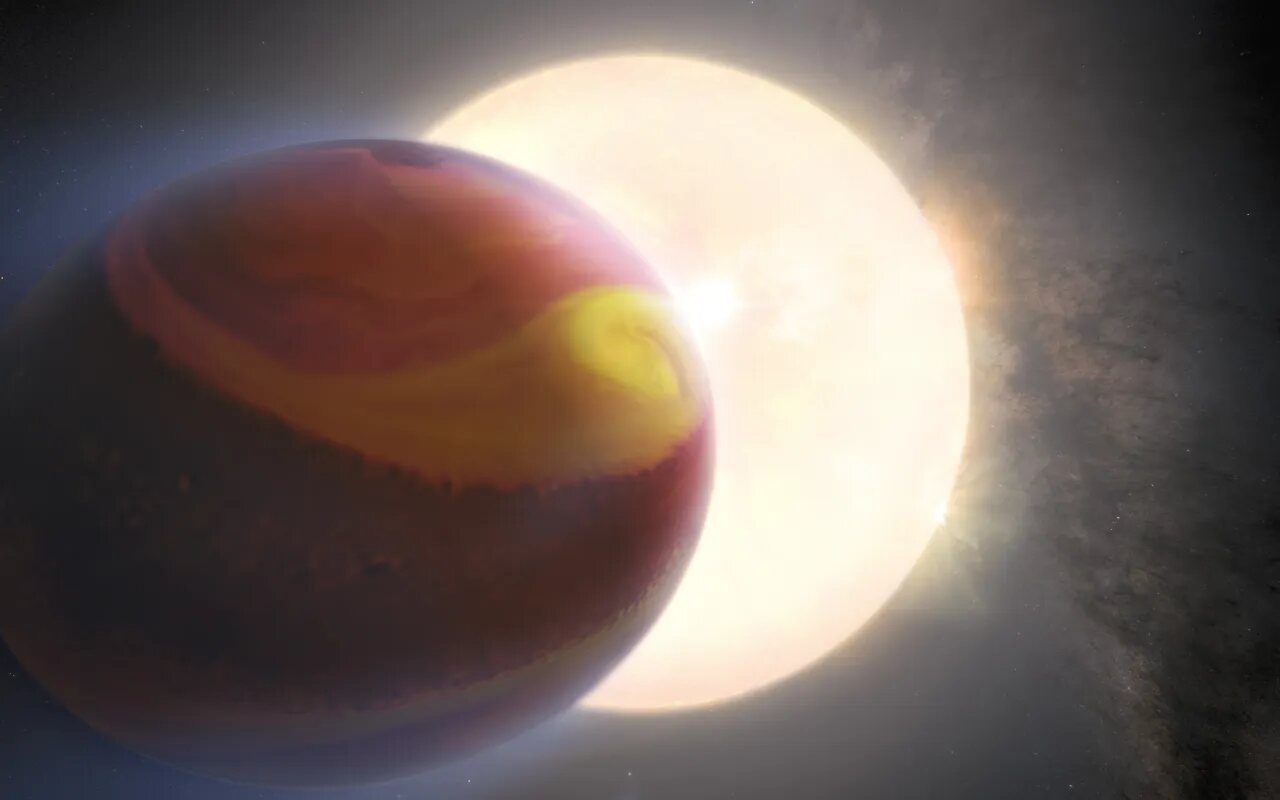
Located a staggering 880 light-years away, WASP-121 b is a massive Jupiter-sized planet that has captivated the attention of scientists. By combining several years of Hubble observations with sophisticated supercomputer modeling, astronomers have generated stunning evidence for the presence of massive cyclones and other dynamic weather activities on this fiery exoplanet.
Just like our own solar system, neighboring planets exhibit ever-changing atmospheric conditions. However, unraveling the complexities of exoplanet weather patterns requires an immense amount of detailed observations and cutting-edge computational techniques. Through their meticulous analysis, the international team of astronomers discovered that WASP-121 b's atmosphere is far from static - it is a living, breathing entity, constantly evolving over time.
A Window into Ever-Changing Skies:
The team's journey began by reprocessing and analyzing Hubble observations of WASP-121 b taken in 2016, 2018, and 2019. The results were astonishing. Notable differences in the exoplanet's atmospheric composition, accompanied by massive weather fronts, storms, and cyclones, were observed. These weather phenomena were generated and destroyed due to the stark temperature difference between the illuminated side of the planet and the dark side facing away from its star.
The team's findings were not mere observations but a revelation of the intricate dance of nature. By employing sophisticated modeling techniques, they pieced together the puzzle of temporal variations in the exoplanet's atmosphere. Through their simulations, they were able to accurately map the ever-changing weather patterns on ultra-hot planets like WASP-121 b.
Multiple Perspectives in the Quest for Knowledge:
In the pursuit of unraveling the secrets of the universe, collaboration across borders and diverse perspectives is crucial. This extraordinary discovery was made possible by a team of international astronomers, each bringing their unique expertise to the table. From the European Space Agency to the California Institute of Technology, Brandeis University to the University College London, this diverse group united to venture into unknown territories and push the boundaries of our understanding.
Inspiring Future Explorers:
This remarkable achievement is more than just a scientific breakthrough; it ignites the flame of exploration within us all. The tantalizing glimpse into the ever-changing atmosphere of distant exoplanets encourages us to continue pushing the boundaries of discovery. It sparks a fascination for the unknown and fuels our passion for unraveling the mysteries of the cosmos.
Looking Ahead:
With this groundbreaking research as a guiding light, the possibilities for future investigations and exploration are boundless. As Hubble embarks on its latest cycle of observations, we can only imagine the wonders it will uncover and the previously unseen worlds it will reveal.
Conclusion:
NASA's Hubble Space Telescope continues to amaze us, offering a window into the infiniteness of the universe. Its recent observations of WASP-121 b's evolving exoplanet atmosphere over a period of three years have elevated our understanding of the dynamic nature of distant worlds. It reminds us that the secrets of the universe are waiting to be discovered, and by collaborating across diverse perspectives, we can unlock the mysteries of our cosmic existence. Let us be inspired to explore, to question, and to keep reaching for the stars.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?