A recent study by the National Institutes of Health found that Artificial intelligence (AI) and machine learning (ML) can be used to detect and diagnose Polycystic Ovary Syndrome (PCOS). PCOS is the most common hormone disorder among women between the ages of 15 and 45. The researchers reviewed published scientific studies that used AI/ML to analyze data to diagnose and classify PCOS. They concluded that AI/ML-based programs were successful in detecting PCOS.
“Given the large burden of under- and misdiagnosed PCOS in the community and its potentially serious outcomes, we wanted to identify the utility of AI/ML in the identification of patients that may be at risk for PCOS,” said Janet Hall, M.D., senior investigator and endocrinologist at the National Institute of Environmental Health Sciences (NIEHS), part of NIH, and a study co-author. “The effectiveness of AI and machine learning in detecting PCOS was even more impressive than we had thought.”
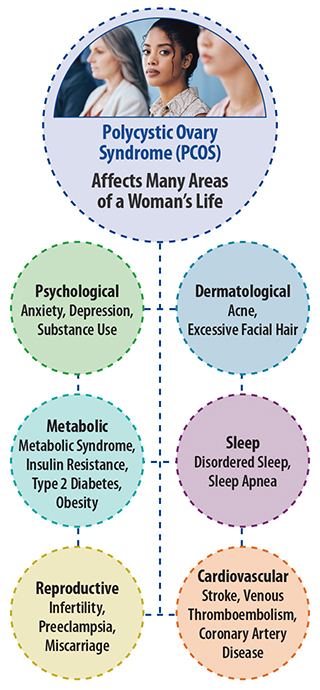
Polycystic ovary syndrome (PCOS) is a hormonal disorder that affects the proper functioning of the ovaries. In many cases, it is accompanied by higher levels of testosterone. This condition can lead to a range of symptoms, including irregular periods, acne, excessive facial hair growth, or baldness. Women with PCOS are more likely to develop type 2 diabetes, as well as sleep, psychological, cardiovascular, and other reproductive disorders such as uterine cancer and infertility.
“PCOS can be challenging to diagnose given its overlap with other conditions,” said Skand Shekhar, M.D., senior author of the study and assistant research physician and endocrinologist at the NIEHS.
“These data reflect the untapped potential of incorporating AI/ML in electronic health records and other clinical settings to improve the diagnosis and care of women with PCOS,” said Shekhar.
The diagnosis of polycystic ovary syndrome (PCOS) is based on standardized criteria that include clinical features, laboratory findings, and radiological evidence. However, PCOS is often difficult to diagnose because some of its symptoms can overlap with other conditions. To improve the accuracy of diagnosis, researchers suggest integrating large population-based studies with electronic health datasets and using machine learning (ML) to identify sensitive diagnostic biomarkers. ML is a type of artificial intelligence (AI) that can process large amounts of data, such as electronic health records.
In a recent study, researchers conducted a systematic review of peer-reviewed studies published in the last 25 years that used AI/ML to diagnose PCOS. They screened 135 studies and included 31 in their analysis. The studies were all observational and assessed the use of AI/ML technologies for diagnosing PCOS. Ultrasound images were used in about half of the studies, and the average age of the participants was 29.
The accuracy of PCOS diagnosis using AI/ML varied across the studies. Among the 10 studies that used standardized diagnostic criteria, the accuracy of detection ranged from 80-90%. The findings suggest that AI/ML can be a valuable tool for diagnosing difficult-to-diagnose disorders like PCOS.
“Across a range of diagnostic and classification modalities, there was an extremely high performance of AI/ML in detecting PCOS, which is the most important takeaway of our study,” said Shekhar. The use of AI/ML-based programs has the potential to significantly improve our ability to detect women with PCOS early, which can lead to cost savings and lessen the burden of PCOS on patients and the health system. Further studies with strong validation and testing practices will enable a smooth integration of AI/ML for chronic health conditions. NIEHS clinical studies are currently underway to understand and detect PCOS. To learn more, you can join an NIEHS study.
The findings of the NIH study are clear: AI/ML can be used to successfully diagnose Polycystic Ovary Syndrome. This is a major breakthrough in the medical field, as it demonstrates the potential of AI/ML to improve the accuracy and speed of diagnosis for this and other conditions. With further research and development, AI/ML could revolutionize the way medical professionals diagnose and treat patients.
Grants: This work was supported by the Intramural Research Program of the NIH/National Institute of Environmental Health Sciences (ZIDES102465 and ZIDES103323).


 How to resolve AdBlock issue?
How to resolve AdBlock issue?