Chemists from RUDN University, located in Moscow, along with their colleagues in China, have successfully developed several machine-learning models to identify potential drugs that can restrict the activity of an enzyme, Cyclin-dependent kinase 2 (CDK 2), which is responsible for uncontrolled cell division. Although CDK 2 is not necessary for healthy cells, it plays a crucial role in the uncontrolled growth of cancer cells. Inhibiting the activity of CDK 2 can restrain tumor growth, making it crucial to find effective CDK 2 inhibitors. The chemists from RUDN University and their colleagues in China used a combination of machine learning and molecular modeling techniques to identify several potential inhibitors.
"Cyclin-dependent kinase 2 is a promising target for cancer treatment. The development of its inhibitors is important in antitumor therapy. The participation of this enzyme in tumor formation remains incompletely studied, but it is already clear that its inhibition is useful in the treatment of cancer. Several inhibitors have already undergone clinical trials, but a selective inhibitor specifically for this enzyme has not yet been found," said Alexander Novikov, Ph.D. in Chemistry, senior researcher at the Joint Institute of Chemical Research of RUDN University.
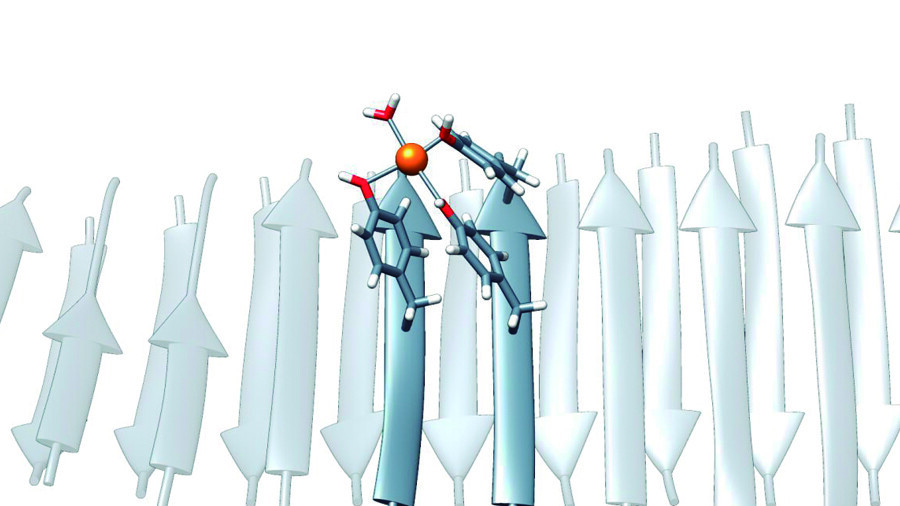
Chemists utilized machine learning methods to identify a potential candidate drug. The authors of the study developed multiple models to find active inhibitors of CDK 2. Additionally, they built a molecular model using the molecular docking method, which can identify the most favorable molecular orientation for forming a stable complex.
Using machine learning models, the team identified 25 potential active CDK 2 inhibitors with an accuracy of 98%. Chemists then tested each of the identified inhibitors using molecular docking. Out of the 25, three substances proved to work better than the rest. For the top three, a computer simulation was created using the molecular dynamics method and compared with the reference compound, dalpiciclib. The results showed that all three of the chosen inhibitors were more stable and more compact than the reference compound.
"Compared to the control drug dalpiciclib, the three calculated compounds showed more stable behavior and compactness. Despite the promising results, our study has several limitations. We need in-depth clinical trials in vitro and in vivo to confirm inhibitory activity and potential therapeutic efficacy. In addition, when developing drugs, it will be necessary to study the effect of compounds on off-target interactions and their toxicity," Alexander Novikov, Ph.D. in Chemistry, senior researcher at the Joint Institute of Chemical Research of RUDN University.
The research concludes that Russian chemists have utilized machine learning and molecular modeling effectively to detect new potential anticancer drugs. This is a significant breakthrough in cancer research that could save countless lives by leading to the development of new treatments. The research also exemplifies the power of combining modern technology with traditional scientific methods, demonstrating the potential of machine learning and molecular modeling to revolutionize the field of medicine.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?