In cereal crops, the number of new leaves each plant produces is used to study the periodic events that constitute the biological life cycle of the crop. The conventional method of determining leaf numbers involves manual counting, which is slow, labor-intensive, and usually associated with large uncertainties because of the small sample sizes involved. It is thus difficult to get accurate estimates of some traits by manually counting leaves.
Conventional methods have, however, been improved upon with technology. Deep learning has enabled the use of object detection and segmentation algorithms to estimate the number of plants (and leaves on these plants) in an area. There is, however, a roadblock to using these algorithms. They count leaf tips, which appear tiny in images, proving difficult to detect. Consequently, deep learning methods often fail to perform in actual field conditions. Aiming to solve this problem, a multinational research team developed a self-supervised leaf-tip counting method based on deep learning techniques, which yielded wheat leaf count with high accuracy. The study was led by Professor Shouyang Liu of the Nanjing Agricultural University located in Nanjing, Jiangsu Province, China. and was published online in Plant Phenomics on March 20, 2023.
Speaking about their work, Prof. Liu says, “We developed a high-throughput method to count the number of leaves on wheat plants by detecting leaf tips in RGB (red-green-blue) images. The Digital Plant Phenotyping platform (D3P) was used to simulate a large, diverse dataset of RGB images and corresponding leaf-tip labels of wheat plant seedlings. Over 150,000 images were generated, with over 2 million labels.”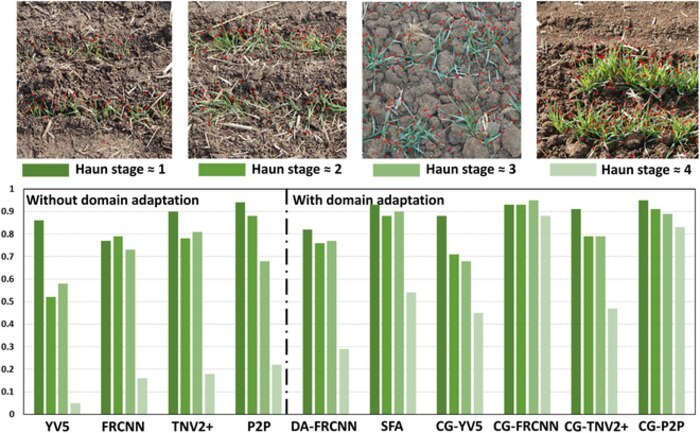 
The researchers used domain adaptation—in which a neural network trained on a “source” dataset is applied to a “test” dataset, also referred to as a “target” dataset. This was achieved through deep learning techniques that mimic neural processes used by the human brain and use algorithms inspired by its structure and function.
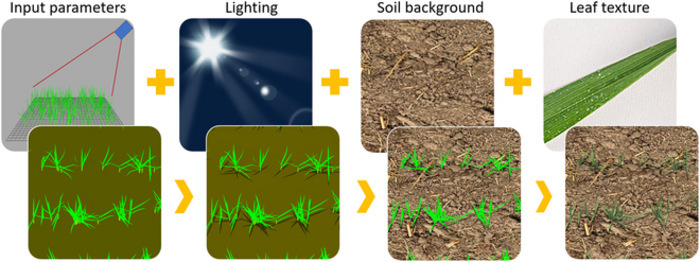
Next, the researchers collected 2,763 RGB images of juvenile wheat fields from 11 locations spread across five countries. A variety of measures were used to create a robust and reliable source dataset—different types of cameras, varying imaging angles, and images with diverse soil backgrounds/light conditions were used. Besides capturing field images, the team also generated simulated wheat images, which were automatically annotated using the D3P. Domain adaptation was used to improve the realism of these images, which were then used to train the deep-learning models.
Six combinations of deep learning models and domain adaptation techniques were used in this study; the Faster-RCNN model with the CycleGAN adaptation technique demonstrated the best performance. This was evident from its high coefficient of determination (R2 = 0.94)—a measure that determines the goodness of fit of a model—and optimal root mean square error (RMSE = 8.7)—a standard way to measure the error of a model in predicting quantitative data.
Moreover, of the three factors evaluated for the performance of the leaf counting models, the light condition was found to be of utmost importance. On the other hand, leaf texture and soil brightness were found to be less important for performance, but the combination of all three factors was found to significantly improve the realism of the images. The results also revealed that a spatial resolution higher than 0.6 mm per pixel was required to ensure accurate identification of leaf tips.
Explaining the implications of their study, Prof. Liu says, “The resulting proposed deep learning method appears very attractive since it eliminates the tedious, expensive, and sometimes inaccurate manual labeling task by simulating images for which the labels are automatically generated. The images were also made more realistic using domain adaptation techniques.”
The research team has made the trained networks available at https://github.com/YinglunLi/Wheat-leaf-tip-detection to facilitate further research in this area.


 How to resolve AdBlock issue?
How to resolve AdBlock issue?