Yellow fever is a deadly disease in overpopulated tropical regions of Africa and South America. Infected people have a temperature increase to 39-41oC, chills, severe headache, nausea, and vomiting. The patient’s face becomes dull, the eyelids swell and the skin turns yellow due to liver damage (hence the name of the disease). Before the yellow fever vaccine was developed, the infection claimed thousands of lives for example in 1871, 8 percent of the population of Buenos Aires died in the epidemic. In mosquito-infested areas, where the vaccination is not readily available to the majority of the population, outbreaks of infection still occur. The yellow fever virus, as well as its related flaviviruses causing Zika and Dengue fever, is treated only by symptomatic treatment, as there are no specific drugs. An international team of scientists used artificial intelligence to select from a vast array of molecules that might be suitable for this purpose. Scientists from the Research Centre of Biotechnology of the Russian Academy of Sciences developed the technology and purchased or synthesized five of the most promising compounds and investigated their activity. The research was conducted in cooperation with Collaborations Pharmaceuticals, Inc. a private company specializing in innovative therapeutics for multiple rare and infectious diseases (based in the USA), São Carlos Institute of Physics, University of São Paulo (Brazil) along with support from the NIH, NIAID (USA). 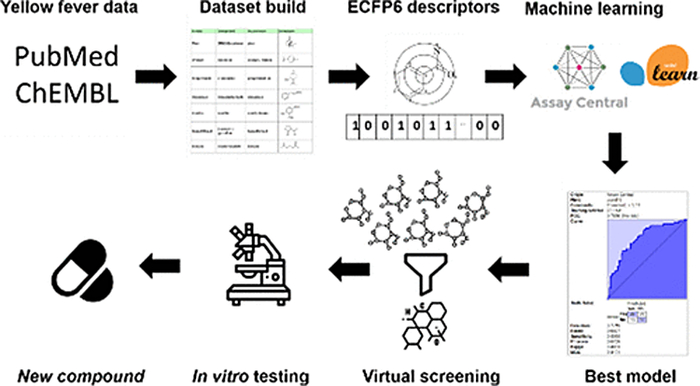
“Our team used a predictive computer model in combination with several machine learning methods. For model training, we relied on in vitro screening data and information available in existing databases to select identify the ideal molecule features for desired activity. With the help of these computational models we predicted their bioactivity before testing them in vitro using NIAID resources”, — explains Vadim Makarov, the co-author, Dr.Sci. (Pharmacy), the head of the Laboratory for Biomedicinal Chemistry of Research Centre of Biotechnology RAS.
Typically, only one of the 5,000 molecules that survived experimental testing is given a chance to reach the pharmacy counter. Others are too toxic, hard to produce, disintegrate in the body, or show too little activity in the real body compared to the test tube. Selection is even more rigorous before the experiments. Even if you focus on the hundreds of thousands of molecules that are known to science that are used or used to treat something else, testing them all not the same on animals and humans, but even in vitro would be almost infinite. To make the first stages of experiments cheaper and faster, scientists use supercomputer simulations and try to convert some of the initial tests into virtual ones. In the next stage, they are also assisted by high-throughput screening, during which “the robot dispenser” automatically dispenses tiny amounts of active substances into the microplates that contain cells infected with viruses. The researcher then evaluates which compounds kill the virus.
The authors of the paper created supercomputer models that can self-learn, comparing chemical compounds according to certain structural rules. Machine learning requires as much basic information from molecules wit or without activity as possible. For this purpose, scientists took information from public databases on small medicinal molecules and studied scientific publications on yellow fever virus research on cells. The models helped propose five of the most promising molecules that would fight the virus in human cells. Scientists have then tested these molecules and found the optimal concentration at which they should work. For the most efficient substance, the half-maximal effective concentration was 3.2 uM (equal to one mole of active substance per liter).
“The molecule we choose relates to the derivatives of pyrazosulphonamide. Its activity with the yellow fever virus is so great that we can talk about a potential drug. The structure of this molecule provides ample opportunity for further modification, which could greatly expand the list of potentially affordable yellow fever drugs. If the tests are successful, we will receive an entirely new group of drugs to fight this dangerous disease”, — says Vadim Makarov.


 How to resolve AdBlock issue?
How to resolve AdBlock issue? 
