When a person sneezes or coughs, they can potentially transmit droplets carrying viruses like SARS-CoV-2 to others in their vicinity. Does talking to an infected person also carry an increased risk of infection? How do speech droplets or “aerosols” move in the air space between the people interacting? 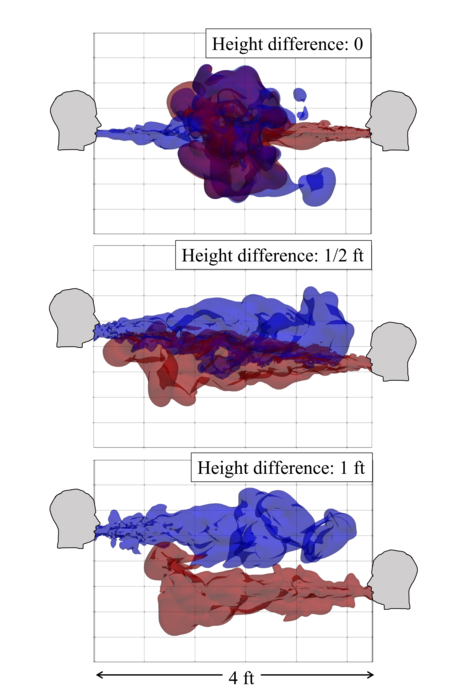
To answer these questions, a research team has carried out supercomputer simulations to analyze the movement of speech aerosols. The team includes researchers from the Department of Aerospace Engineering, Indian Institute of Science (IISc), along with collaborators from the Nordic Institute for Theoretical Physics (NORDITA) in Stockholm and the International Centre for Theoretical Sciences (ICTS) in Bengaluru. Their study was published in the journal Flow.
The team visualized scenarios in which two maskless people are standing two, four, or six feet apart and talking to each other for about a minute, and then estimated the rate and extent of spread of the speech aerosols from one to another. Their simulations showed that the risk of getting infected was higher when one person acted as a passive listener and didn’t engage in a two-way conversation. Factors like the height difference between the people talking and the number of aerosols released from their mouths also appear to play an important role in viral transmission.
“Speaking is a complex activity … and when people speak, they’re not really conscious of whether this can constitute a means of virus transmission,” says Sourabh Diwan, Assistant Professor in the Department of Aerospace Engineering, and one of the corresponding authors.
In the early days of the COVID-19 pandemic, experts believed that the virus mostly spread symptomatically through coughing or sneezing. Soon, it became clear that asymptomatic transmission also leads to the spread of COVID-19. However, very few studies have looked at aerosol transport by speech as a possible mode of asymptomatic transmission, according to Diwan.
To analyze speech flows, he and his team modified a computer code they had originally developed to study the movement and behavior of cumulus clouds – the puffy cotton-like clouds that are usually seen on a sunny day. The code (called Megha-5) was written by S Ravichandran from NORDITA, the other corresponding author on the paper, and was used recently for studying particle-flow interaction in Rama Govindarajan’s group at ICTS. The analysis carried out by the team on speech flows incorporated the possibility of viral entry through the eyes and mouth in determining the risk of infection – most previous studies had only considered the nose as the point of entry.
“The computational part was intensive, and it took a lot of time to perform these simulations,” explains Rohit Singhal, first author and Ph.D. student at the Department of Aerospace Engineering. Diwan adds that it is hard to numerically simulate the flow of speech aerosols because of the highly fluctuating (“turbulent”) nature of the flow; factors like the flow rate at the mouth and the duration of speech also play a role in shaping its evolution.
In the simulations, when the speakers were either of the same height or drastically different heights (one tall and another short), the risk of infection was found to be much lower than when the height difference was moderate – the variation looked like a bell curve. Based on their results, the team suggests that just turning their heads away by about nine degrees from each other while still maintaining eye contact can reduce the risk for the speakers considerably.
Moving forward, the team plans to focus on simulating differences in the loudness of the speakers’ voices and the presence of ventilation sources in their vicinity to see what effect they can have on viral transmission. They also plan to engage in discussions with public health policymakers and epidemiologists to develop suitable guidelines. “Whatever precautions we can take while we come back to normalcy in our daily interactions with other people, would go a long way in minimizing the spread of infection,” Diwan says.


 How to resolve AdBlock issue?
How to resolve AdBlock issue? 
