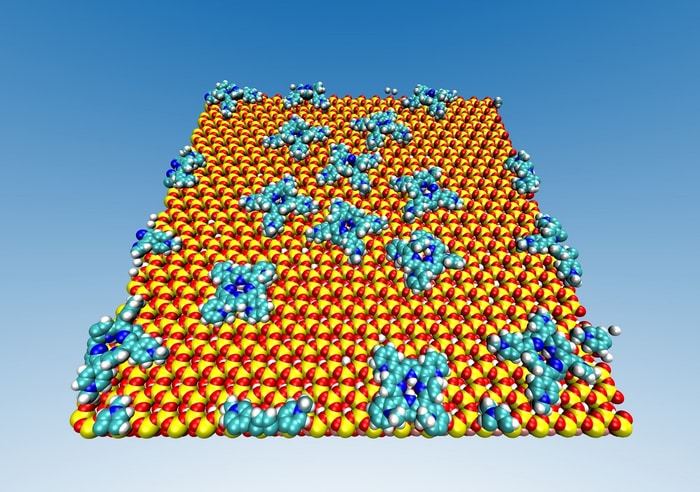
Symmetry is a prevalent feature of nature at all scales. For example, our naked eyes can easily identify symmetries in the bodily shape of countless organisms. Symmetry is also very important in the fields of physics and chemistry, especially in the microscopic realm of atoms and molecules. Crystals, which are highly ordered materials, can even have multiple types of symmetry at the same time, such as rotational symmetry, inversion symmetry, and translational symmetry.
Lately, alongside rapid progress in computer science, researchers have developed computational methods that seek to predict the physical properties of crystals based on their electronic structure. In practice, however, pure and perfectly symmetric crystals are seldom used. This is because a crystal’s properties can be tuned as desired by alloying them with other materials or randomly substituting certain atoms with other elements, i.e., doping.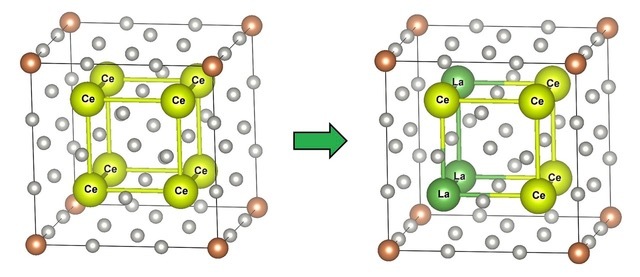
Accordingly, materials scientists are seeking computationally efficient approaches to analyze such alloys and substituted crystals, also known as solid solutions. The ‘supercell method’ is one such approach and is widely used to model crystal structures with random substitutions of different atoms. The symmetry of crystals, however, is actually a problem when using this technique. In crystals, there can be many substitution patterns that are physically equivalent to other substitutions if we simply translate or rotate them. Findings of these symmetric substitution patterns are not very meaningful, and thus their calculation when using the supercell method is a waste of time.
In a recent study, a team of researchers led by Assistant Professor Kousuke Nakano from the Japan Advanced Institute of Science and Technology (JAIST) found a solution to this problem. They developed an open-source software called “Suite for High-throughput generation of models with atomic substitutions implemented by Python,” or SHRY that can, in terms of symmetry, generate distinct substitution patterns in solid solutions and alloys [https://github.com/giprayogo/SHRY]. This work, which was published in the ACS Journal of Chemical Information and Modeling, was co-authored by doctoral student Genki I. Prayogo, Dr. Andrea Tirelli, Professor Ryo Maezono, and Associate Professor Kenta Hongo.
The team approached the problem from the angle of group theory. It turns out that searching for atomic substitution patterns in crystals is analogous to the problem of finding coloring patterns on the vertices of graphs under certain restrictions. This allows one to reformulate the original problem of finding non-symmetric atomic substitutions in crystals as exploring search trees depicting the coloring of vertices in graphs.![Figure 2. Example of a search tree representing an atomic substitution CAPTION: In this search tree, three vertices out of eight are colored red in a cubic arrangement, representing the substitution of Figure 1. p(X) denotes the parent node of X, whereas C(X) denotes the set of children of X. Because of the symmetry of the cube, [p(X) and p(Y)], [X and Y] and [Z and W] are identical. Image credit: Kousuke Nakano from JAIST. Figure 2. Example of a search tree representing an atomic substitution CAPTION: In this search tree, three vertices out of eight are colored red in a cubic arrangement, representing the substitution of Figure 1. p(X) denotes the parent node of X, whereas C(X) denotes the set of children of X. Because of the symmetry of the cube, [p(X) and p(Y)], [X and Y] and [Z and W] are identical. Image credit: Kousuke Nakano from JAIST.](/images/igure2treeexample_ab418.jpeg) However, the way in which the search tree is explored is crucial. A simple, naïve approach in which all possible branches are searched and directly compared is impossible; the time and calculations required to grow uncontrollably for large systems. This happens because deciding whether to explore further down a branch requires information about all other branches besides the one being explored, which is technically referred to as ‘non-local information’.
However, the way in which the search tree is explored is crucial. A simple, naïve approach in which all possible branches are searched and directly compared is impossible; the time and calculations required to grow uncontrollably for large systems. This happens because deciding whether to explore further down a branch requires information about all other branches besides the one being explored, which is technically referred to as ‘non-local information’.
To avoid this issue, the researchers implemented in SHRY a technique called canonical augmentation. “This method can decide whether a tree branch should be explored more deeply or not based solely on local information,” explains Dr. Nakano, “Most importantly, theorems from group theory guarantee that only distinct substitution patterns will be extracted, without over-or under-exploring the tree structure in the terms of symmetry.” The team verified that their algorithm was error-free by testing it thoroughly with data from a database of crystal structures.
It is worth noting that SHRY was written in Python 3, one of the most popular cross-platform programming languages, and uploaded to GitHub, a leading project-sharing online platform. “SHRY can be used as a stand-alone program or imported into another Python program as a module,” highlights Dr. Nakano, “Our software also uses the widely supported Crystallographic Information File (CIF) format for both the input and output of the sets of substituted crystal structures.” The team plans to keep improving SHRY’s code based on feedback from other users, boosting its speed and capabilities.
Overall, the software developed in this study could help scientists identify potential atomic substitutions in solids, which is the most common strategy used to tune the properties of materials for practical applications. SHRY will help speed up research and develop substituted crystals with unprecedented functionalities and superior characteristics.
Dr. Donald C. Wunsch II, the Mary K. Finley Missouri Distinguished Professor of Electrical and Computer Engineering at Missouri University of Science and Technology, has been appointed founding director of the university’s Kummer Institute Center for Artificial Intelligence and Autonomous Systems. Wunsch will begin his duties on Dec. 1, when he completes his appointment as a program director in the National Science Foundation’s Energy, Power, Control and Networks program. 
As founding director, Wunsch will lead the development of the new center’s mission, vision, and strategic plan to develop a nationally recognized research center focused on the rapidly evolving fields of artificial intelligence and autonomous systems. Ultimately, the center will deploy the latest machine learning and artificial intelligence techniques to advance the performance and intelligence of a range of robotic and autonomous solutions. Wunsch will also be responsible for leveraging the breadth of capabilities and expertise among S&T’s faculty to strengthen collaboration with and among several university-based research centers.
The center was founded to spur regional, national and global development in artificial intelligence and autonomous systems, in alignment with the vision of the Kummer Institute for Student Success, Research and Economic Development at Missouri S&T. Late St. Louis entrepreneur Fred Kummer, a Missouri S&T graduate, and his wife June donated $300 million to Missouri S&T in October 2020 to establish the Kummer Institute. Their donation is the largest single gift ever to any public or private university in Missouri and one of the largest to any university.
“Mr. Kummer recognized the potential artificial intelligence and autonomous systems have to truly change the world,” says Dr. Kamal Khayat, interim vice chancellor for research and innovation at S&T. “I am pleased to welcome someone as accomplished and capable as Dr. Wunsch to lead our institute’s innovative efforts in these two truly transformative disciplines. The insights and expertise he has acquired throughout his career will be an asset to the entire S&T community as he steps into this new role.”
Before joining S&T in 1999, from 1984 to 1993, Wunsch worked for Boeing, culminating in his role as a senior principal scientist. In 1993 he joined Texas Tech University as an assistant professor with a joint appointment to the departments of electrical and computer engineering and computer science; he was promoted to associate professor at Texas Tech in 1998. He joined the electrical and computer engineering department at S&T in 1999 and has courtesy appointments in system engineering, computer science, business administration and mathematics, and statistics. In 2005, Wunsch served as president of the International Neural Networks Society (INNS), which bestowed upon him its Gabor Award in 2015 and its Ada Lovelace Award in 2019. He is a Fellow of the INNS and the Institute of Electrical and Electronics Engineers (IEEE) and was twice the Charles Hedlund Distinguished Visiting Professor at the American University in Cairo. Wunsch has mentored 23 Ph.D. recipients in computer engineering, electrical engineering, systems engineering, and computer science.
Wunsch’s research has focused on unsupervised learning, also known as clustering, particularly hierarchical and real-time versions; adaptive resonance and reinforcement learning architectures; hardware and applications; and various multidisciplinary collaborations. He has more than 500 publications to his credit, including 12 books.
“I feel honored and grateful for the phenomenal opportunity to help develop the Kummer Institute Center for Artificial Intelligence and Autonomous Systems,” Wunsch says. “For over two decades, I’ve been amazed at the novelty, diversity, and impact of S&T research. I look forward to supporting collaborations across campus, across our region, and around the world as we grow and intensify our competitiveness in these incredibly transformational fields. I would like to hear from internal and external stakeholders about how AI and Autonomous Systems can help them achieve their goals.”
Wunsch earned a doctorate in electrical engineering and a master’s degree in applied mathematics from the University of Washington in 1991 and 1987, respectively. He also earned an executive M.B.A. from Washington University in 2006 and a bachelor’s degree in applied mathematics from the University of New Mexico in 1984 and completed the Jesuit Core Honors Program at Seattle University in 1981.
The Center for Artificial Intelligence and Autonomous Systems is one of four new research centers established through the Kummer Institute. The Center for Advanced Manufacturing is led by Dr. Richard Billo, who joined S&T in January. National searches for founding directors continue for the Center for Advanced and Resilient Infrastructure and the Center for Resource Sustainability.


 How to resolve AdBlock issue?
How to resolve AdBlock issue? 