Six months ago, astronomers at the University of Bonn reported the discovery of an extremely long intergalactic gas filament with the X-ray telescope eROSITA. A new study has now focused on an interesting structure in the filament, the northern clump. Their new observational data prove that this is a cluster of galaxies with a black hole at its center. Therefore, the gas filament is a galactic matter highway: The northern clump is moving along it towards two more giant galaxy clusters and will eventually merge with them. The paper will be published in the journal Astronomy & Astrophysics, along with other papers published on the occasion of the first eROSITA data release.
The universe resembles a Swiss cheese - but one with huge holes: Large areas in space are absolutely empty. In between, thousands of galaxies crowd in a comparatively small space. These clusters are connected by highways of thin matter gas, like the gossamer filaments of a spider's web. 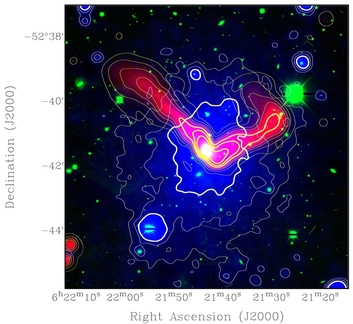
At least, this is what the standard model of cosmology predicts. Whether this is actually the case was hard to prove until recently. This is because the matter in the gas filaments is so diluted that it eluded the view of even the most sensitive measuring instruments: The filaments contain just ten particles per cubic meter, which is far fewer than are present in the best vacuum that humans can produce.
This is why a study led by the University of Bonn in Germany caused such a stir last winter. The researchers had discovered an intergalactic gas filament measuring at least 50 million light-years in length that emanates from two giant galaxy clusters. "There is another galaxy cluster in this filament, the northern clump," explains Prof. Dr. Thomas Reiprich of the Argelander Institute for Astronomy at the University of Bonn. "In the paper now submitted for publication, we have taken a closer look at this."
Bow shock and matter tail
To do this, the researchers combined images from several sources: the SRG/eROSITA, XMM-Newton, and Chandra satellites, as well as the EMU survey with the ASKAP radio telescope and DECam optical data. The resulting images have a richness of detail never seen before. "This allows us to identify a large galaxy at the center of the northern clump," says Reiprich's colleague and lead author of the study, Angie Veronica. "And at its center sits a supermassive black hole." Two so-called matter jets emanate from it, in which the particles move away from the black hole at close to the speed of light. This produces synchrotron radiation, which can be visualized in radio telescope images.
In addition, the northern clump contains very hot matter gas. "Because of its high temperature of 20 million degrees, it emits X-rays, which we see in the eROSITA images and have now been able to measure very precisely with the XMM-Newton satellite," says Veronica. Overall, the combination of data sources indicates that the northern clump is likely moving at high velocity. The jets of matter emanating from the black hole point backward like the braids of a running girl; in front of the clump, the gas additionally seems to form a kind of bow shock. "We also see a matter tail behind it," Reiprich explains. "We currently interpret this observation to mean that the northern clump is losing matter as it travels. However, it could also be the case that even smaller clumps of matter in the highway are falling towards the northern clump."
Overall, the observations confirm the view derived from theories that the gas filament is an intergalactic matter highway. The northern clump is moving along this road at high speed toward two other much larger clusters of galaxies called Abell 3391 and Abell 3395. "It falls on these piles, so to speak, and will continue to make them bigger - according to the principle: Whoever has will be given more," explains Reiprich, who is also a member of the transdisciplinary research area "Building Blocks of Matter" at the University of Bonn. "What we're seeing is a snapshot of this fall."
Observations consistent with theoretical predictions
The observations are remarkably consistent with the result of the Magneticum supercomputer simulations developed by researchers of the eROSITA consortium. Therefore, they can also be taken as an argument that the current assumptions about the origin and evolution of the universe are correct. This includes the thesis that a large part of the matter is invisible to our measuring instruments. 85 percent of the matter in our universe is said to consist of this "dark matter". One of its most important roles in the standard model of cosmology is as a condensation nucleus, which caused gaseous matter to condense into galaxies after the Big Bang.


 How to resolve AdBlock issue?
How to resolve AdBlock issue? 

