Researchers have discovered that light, in the form of a laser, can trigger a form of magnetism in a normally nonmagnetic material. This magnetism centers on the behavior of electrons. These subatomic particles have an electronic property called “spin,” which has a potential application in quantum supercomputing. The researchers found that electrons within the material became oriented in the same direction when illuminated by photons from a laser.
The experiment was led by scientists at the University of Washington and the University of Hong Kong. 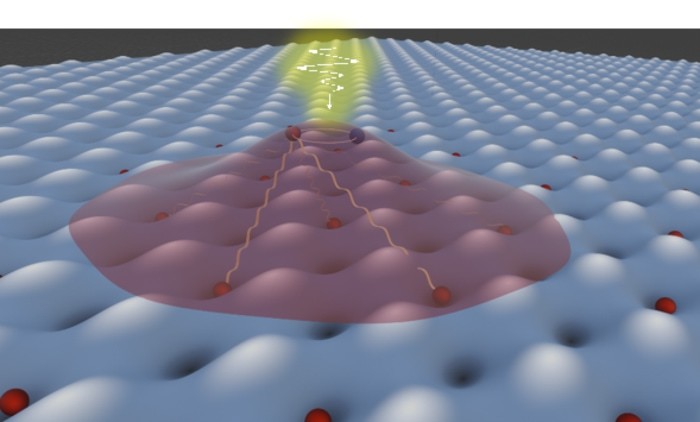
By controlling and aligning electron spins at this level of detail and accuracy, this platform could have applications in the field of quantum simulation, according to co-senior author Xiaodong Xu, a Boeing Distinguished Professor at the UW in the Department of Physics and the Department of Materials Science and Engineering.
“In this system, we can use photons essentially to control the ‘ground state’ properties — such as magnetism — of charges trapped within the semiconductor material,” said Xu, who is also a faculty researcher with the UW’s Clean Energy Institute and the Molecular Engineering & Sciences Institute. “This is a necessary level of control for developing certain types of qubits — or ‘quantum bits’ — for quantum computing and other applications.”
Xu, whose research team spearheaded the experiments, led the study with co-senior author Wang Yao, professor of physics at the University of Hong Kong, whose team worked on the theory underpinning the results. Other UW faculty members involved in this study are co-authors Di Xiao, a UW professor of physics and materials science and engineering who also holds a joint appointment at the Pacific Northwest National Laboratory, and Daniel Gamelin, a UW professor of chemistry and director of the Molecular Engineering Materials Center.
The team worked with ultrathin sheets — each just three layers of atoms thick — of tungsten diselenide and tungsten disulfide. Both are semiconductor materials, so named because electrons move through them at a rate between that of a fully conducting metal and an insulator, with potential uses in photonics and solar cells. Researchers stacked the two sheets to form a “moiré superlattice,” a stacked structure made up of repeating units.
Stacked sheets like these are powerful platforms for quantum physics and materials research because the superlattice structure can hold excitons in place. Excitons are bound pairs of “excited” electrons and their associated positive charges, and scientists can measure how their properties and behavior change in different superlattice configurations.
The researchers were studying the exciton properties within the material when they made the surprising discovery that light triggers a key magnetic property within the normally nonmagnetic material. Photons provided by the laser “excited” excitons within the laser beam’s path, and these excitons induced a type of long-range correlation among other electrons, with their spins all orienting in the same direction.
“It’s as if the excitons within the superlattice had started to ‘talk’ to spatially separated electrons,” said Xu. “Then, via excitons, the electrons established exchange interactions, forming what’s known as an ‘ordered state’ with aligned spins.”
The spin alignment that the researchers witnessed within the superlattice is a characteristic of ferromagnetism, the form of magnetism intrinsic to materials like iron. It is normally absent from tungsten diselenide and tungsten disulfide. Each repeating unit within the moiré superlattice is essentially acting as a quantum dot to “trap” an electron spin, said Xu. Trapped electron spins that can “talk” to each other, as these can, have been suggested as the basis for a type of qubit, the basic unit for quantum computers that could harness the unique properties of quantum mechanics for computation.
In a separate paper published Nov. 25 in Science, Xu and his collaborators found new magnetic properties in moiré superlattices formed by ultrathin sheets of chromium triiodide. Unlike the tungsten diselenide and tungsten disulfide, chromium triiodide harbors intrinsic magnetic properties, even as a single atomic sheet. Stacked chromium triiodide layers formed alternating magnetic domains: ferromagnetic one — with spins all aligned in the same direction — and another that is “antiferromagnetic,” where spins point in opposite directions between adjacent layers of the superlattice and essentially “cancel each other out,” according to Xu. That discovery also illuminates relationships between a material’s structure and its magnetism that could propel future advances in supercomputing, data storage, and other fields.
“It shows you the magnetic ‘surprises’ that can be hiding within moiré superlattices formed by 2D quantum materials,” said Xu. “You can never be sure what you’ll find unless you look.”


 How to resolve AdBlock issue?
How to resolve AdBlock issue?